Abstract
Background: To study whether there is genetic overlap underlying the risk for schizophrenia spectrum disorders (SSDs) and low intelligence quotient (IQ), we reviewed and summarized the evidence on genetic variants associated with both traits.
Methods: We performed this review in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) and preregistered it in PROSPERO. We searched the Medline databases via PubMed, PsycInfo, Web of Science and Scopus. We included studies in adults with a diagnosis of SSD that explored genetic variants (single nucleotide polymorphisms [SNPs], copy number variants [CNVs], genomic insertions or genomic deletions), estimated IQ and studied the relationship between genetic variability and both traits (SSD and IQ). We synthesized the results and assessed risk of bias using the Quality of Genetic Association Studies (Q-Genie) tool.
Results: Fifty-five studies met the inclusion criteria (45 case–control, 9 cross-sectional, 1 cohort), of which 55% reported significant associations for genetic variants involved in IQ and SSD. The SNPs more frequently explored through candidate gene studies were in COMT, DTNBP1, BDNF and TCF4. Through genome-wide association studies, 2 SNPs in CHD7 and GATAD2A were associated with IQ in patients with SSD. The studies on CNVs suggested significant associations between structural variants and low IQ in patients with SSD.
Limitations: Overall, primary studies used heterogeneous IQ measurement tools and had small samples. Grey literature was not screened.
Conclusion: Genetic overlap between SSD and IQ supports the neurodevelopmental hypothesis of schizophrenia. Most of the risk polymorphisms identified were in genes relevant to brain development, neural proliferation and differentiation, and synaptic plasticity.
Introduction
Schizophrenia spectrum disorders (SSDs) are characterized by hallucinations, delusions, disorganized thinking, negative symptoms and cognitive dysfunctions that compromise functionality.1,2 They differ from each other according to type of symptoms, duration and etiology; schizophrenia is the most severe and disabling disorder, with a lifetime prevalence of 0.7% to 0.9%.3 The etiology of SSD is unknown, but its onset is influenced by an interaction of genetic and environmental factors.4 Genetic studies have estimated the heritability of schizophrenia to be approximately 80%,5,6 which is explained in part by the global effect of thousands of single nucleotide polymorphisms (SNPs; SNP heritability = 24%).7 In addition, the polygenic burden of risk variants — so-called polygenic risk scores — explains 7.3% of the variance in risk for schizophrenia according to the most recent estimates.7–9
In recent years, there has been widespread interest in establishing endophenotypes of SSD that allow for its identification through observable and quantitative traits.10,11 One of these candidate endophenotypes is IQ,12–16 a quantitative score obtained from a standardized intelligence test that represents an individual’s intellectual ability.17 Thanks to this quantitative estimate of general cognitive function, it is possible to make comparisons between individuals, so it is common to use IQ scores to assess intelligence in the population.17 Current evidence indicates that IQ is heritable, and that its genetic architecture is highly polygenic.18–21 In addition, people with SSD have shown poorer intellectual performance than healthy controls,22,23 findings that in some cases are already present in childhood.24 Along the same line, it has been demonstrated that adolescents at high risk of psychosis exhibit a lower IQ than healthy controls,25 and that the unaffected relatives of patients with SSD present similar deficits.26 Taken together, this evidence indicates that SSDs might be caused by pathological neurodevelopment processes that would be observable as premorbid intellectual deficits.
Genetic overlap between vulnerability to SSD and low intelligence has been identified in large studies involving people with schizophrenia10 and healthy participants.27 This evidence is consistent with the findings of a population-based study in more than 100 000 participants that described significant associations between polygenic risk for schizophrenia and other neuropsychiatric disorders and poorer cognitive functioning, especially in processing speed and memory.28 Accordingly, the large-scale genome-wide association study (GWAS) by Savage and colleagues29 identified 205 genomic loci associated with intelligence, which they found to be enriched in genes expressed in the brain. Moreover, Ohi and colleagues30 found that genetic factors differentiating schizophrenia from bipolar disorder were related specifically to low premorbid intelligence. Thus, the identification of genetic variants that contribute to risk of schizophrenia and low IQ could provide insights into the biological correlates of the disorder.
Generally, different GWASs have found that the genetic variants associated with both traits are enriched in genes expressed in the central nervous system, and that they participate in neurogenesis, regulation of nervous system development, neuronal differentiation and regulation of cell development.7,29,31 For instance, an SNP in the DTNBP1 gene (rs1011313) has been related to both neurocognition and schizophrenia risk, probably because of its involvement in the glutamatergic system.32 Another locus associated with increased risk for schizophrenia and lower general cognition is TCF20 (rs134873, intron variant), which encodes a transcriptional coregulator.33 The authors of that study found that the risk allele of rs134873 was related to increased expression of NAGA (involved in the regulation of glycosylation-associated enzymes, glutamatergic and GABAergic systems34) and decreased expression of CYP2D6 (involved in serotonin and dopamine metabolism) in the human brain.33
Other sources of genetic variation, such as copy number variants (CNVs), may also be involved in the etiology of schizophrenia and intellectual deficits, because patients with schizophrenia who had at least 1 rare CNV yielded low IQs.35
Although several researchers have investigated the genetic variants associated with SSD and IQ, no systematic review has summarized the current findings. We believed that a compilation of results from original studies could contribute in different ways to the field of knowledge. A synthesis with no date limitation allowed us to understand the evolution of an area of research, contributing to a better level of scientific quality and reducing the possibility of bias. Furthermore, by comparing independent samples, a description of positive and negative results helped us to establish whether findings were consistent and could be generalized. Likewise, we wanted to include both candidate gene studies and GWASs to evaluate the replicability of results when using different methods. GWASs yield valuable results, because they have the advantage of reducing possible biases based on their lack of a pre-established hypothesis. However, both GWASs and candidate gene studies are subject to publication bias, because they are at risk of not being reported when the results are negative.
To answer the question of whether overlapping genetic variants underlie the risk of both SSD and low IQ, we aimed to analyze and summarize the findings of primary studies on genetic variants associated with both traits. These data will contribute to establishing IQ as a potential endophenotype for SSD, and to identifying future lines of research.
Methods
This review is reported in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines (PRISMA checklist in Appendix 1, available at www.jpn.ca/lookup/doi/10.1503/jpn.220026/tab-related-content).36 It was registered in the International Prospective Registry of Systematic Reviews (PROSPERO; CRD42020218842), and a protocol was elaborated to guide the review process (https://doi.org/10.21203/rs.3.rs-150210/v1).
Eligibility criteria
We included the following types of studies: published genetic association studies based on IQ (GWASs or candidate gene studies) with an observational design, including cross-sectional, case–control and cohort studies. We considered only manuscripts published in English, but we imposed no restrictions on date of publication. We selected studies in adults with a diagnosis of SSD based on DSM2 or ICD37 criteria at any stage of disease (first episode of psychosis or chronic). We included studies that addressed the association between genetic variability and both traits (SSD and IQ) by exploring the following genetic variants: SNPs, CNVs, genomic insertions and genomic deletions. We included only studies that estimated participants’ IQ using a standardized test. By including studies in patients with SSD that estimated IQ, we sought to guide the search strategy and selection process toward research that targeted the genetic overlap between both traits.
The following were reasons for exclusion from this review: animal model or cell line studies; studies not measuring the outcomes of interest (lacking genotyping, IQ estimation or their association); reviews or meta-analyses; not peer-reviewed; single-case studies; books, editorials or theses.
Search strategies and information sources
We established the design of the search strategy with the advice of an expert librarian from the University of Cantabria. We carried out the search in November 2020 and updated it in October 2021 in the electronic databases of Medline via PubMed, PsycInfo, Web of Science and Scopus.
In accordance with the PRISMA guidelines, we adapted the search strategy to the controlled format of each database when appropriate. For Medline, we used the MeSH format (“Schizophrenia” OR “Psychotic Disorders” OR “Psychosis”) AND (“Genetic Variation” OR “Genetic Variant” OR “Polymorphism, Genetic” OR “Polymorphism, Single Nucleotide” OR “Polymorphism”) AND (“Intelligence” OR “Intelligence Quotient” OR “IQ”). For PsycInfo, we used the Thesaurus format (DE”Schizophrenia” OR “Psychotic Disorders” OR DE”Psychosis”) AND (“Genetic Variation” OR “Genetic Variant” OR “Polymorphism, Genetic” OR “Polymorphism, Single Nucleotide” OR DE”Polymorphism”) AND (DE”Intelligence quotient” OR “IQ”). We screened the Web of Science and Scopus databases using the above terms in free text, and we limited the results in Scopus by document type (article) and language (English) because of the large number of records obtained. We also examined the reference lists of the included articles to identify additional eligible studies.
Study selection and data collection process
After we had retrieved the database results, we recorded them in EndNote (Clarivate Analytics). After duplicate records had been eliminated, 2 reviewers (N.M.G., S.B.M.) screened each record independently by reviewing titles and abstracts. Then, the reviewers separately analyzed the full texts of eligible studies and selected those that met the inclusion criteria, selecting the final studies by consensus. Doubts or discrepancies were resolved between the reviewers, and if necessary, they consulted the postdoctoral researcher (E.S.S.) and the senior researcher (R.A.A.).
Data from the selected studies were recorded in a standardized table designed by the reviewers. The information extracted from each study included the author, the year of publication, the country of origin, the study design, genetic variants and genes investigated, the sample size (patients and healthy controls, if applicable), instruments for measurement of IQ, mean IQ and main findings. Each reviewer collected information from half of the included studies, and then the 2 reviewers exchanged records for verification by their partner.
Outcomes
Among genetic variants for SSD, we considered SNPs, CNVs, genomic insertions and genomic deletions. Only studies that reported the location of the genetic variant were included. For IQ outcomes, we considered a global intelligence score estimated using any standardized measure, such as the Wechsler Adult Intelligence Scale, the Wechsler Test of Adult Reading, the North American Adult Reading Test–Revised (NAART) or the Wechsler Abbreviated Scale of Intelligence.
All types of IQ were included: verbal, performance or full. Verbal IQ is an intelligence index estimated from results on tests of verbal comprehension and working memory; performance IQ is estimated from scores on tests of perceptual organization and processing speed; and full IQ provides a mean of both verbal and performance IQ.17
Quality assessment of primary studies
We used the Quality of Genetic Association Studies (Q-Genie) tool to assess risk of bias in the included studies.38 Q-Genie is a questionnaire of 11 items rated on a 7-point Likert scale; a score of 1 indicates poor quality, and a score of 7 indicates excellent quality. The 11 items evaluate different categories, including the rationale for the study, ascertainment of comparison groups, and technical and nontechnical classification of the genetic variants tested. Q-Genie allowed us to obtain a global score that indicated the overall quality of the study, which could be poor (scores ≤ 35 for studies with control groups and ≤ 32 for studies without control groups), moderate (scores between 35 and 45 for studies with control groups and between 32 and 40 for studies without control groups) or good (scores > 45 for studies with control groups and > 40 for studies without control groups).
Data synthesis
A narrative synthesis of the analyzed studies is presented in the results. For greater clarity and organization of the results, we grouped the findings into 2 sections, 1 on candidate gene studies and 1 on GWASs.
Results
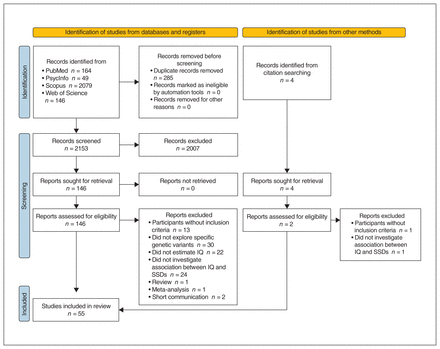
We identified a total of 2438 records from database searching. After duplicates had been eliminated, 2153 results were screened for eligibility, of which 2007 were discarded because they met 1 of the exclusion criteria (3 book chapters, 9 letters to the editor, 24 case studies, 44 meta-analyses, 166 reviews, 226 animal or cellular models, and 1535 did not address the topic of interest). Of the 146 full-text articles we reviewed, 93 were excluded because they did not meet the inclusion criteria. We included 2 additional records from citation searching. In the end, we included 55 articles that explored genetic variants associated with both SSD and IQ in the present review (Figure 1).
PRISMA 2020 flow diagram36 for new systematic reviews which included searches of databases, registers and other sources. PRISMA = Preferred Reporting Items for Systematic Reviews and Meta-Analyses; SSD = schizophrenia spectrum disorder.
Among the included studies, 47 (85.45%) had overall scores of good quality based on the Q-Genie tool, and 8 (14.55%) had scores of overall moderate quality. No study was of low quality (Appendix 1).
Of the 45 studies with a case–control design, 39 (86.67%) reported that patients with SSD had significantly lower IQs than healthy controls, regardless of the evaluation tool used. One of the remaining case–control studies found no significant differences, and the rest (n = 5) did not report participants’ IQ. The included studies explored different polymorphisms, yielding heterogeneous results. However, the genes addressed most frequently using the candidate gene strategy were COMT, BDNF, DTNBP1 and TCF4.
Fewer studies used a GWAS strategy; their findings are presented below.
Candidate gene studies
COMT
Seven studies examined the association of the COMT genotype with both SSD and IQ; 6 had a case–control design and 1 had a cross-sectional design. We found mixed results, as shown in Table 1. Four of the studies (57.14%)39–42 found no significant association between the SNP rs4680 (also known as the Val-158Met polymorphism) and IQ in people with SSD. The other 3 studies (42.86%)43–45 did find significant associations. Green and colleagues44 found that the Val158Met polymorphism was a significant predictor of IQ, because Val homozygotes had lower scores on the intelligence test. Two more studies reported similar results, but with certain specifications. Kontis and colleagues45 found that COMT Val homozygotes had lower IQs, but this effect was reduced by interaction with the rs1801133 allele MTHFR-T. Rebollo-Mesa and colleagues43 observed that the Val158Met polymorphism was significantly related to IQ only in patients who were taking antipsychotics: Val carriers taking a high dose of antipsychotic medication had lower IQs than Met carriers.
Studies exploring the COMT gene and IQ in patients with schizophrenia
BDNF
Five studies explored the link between variations in the BDNF gene and IQ in patients with SSD (Table 2). All studies included a control group and focused on the Val-66Met polymorphism (rs6265). Three studies (60%)46,49,50 reported negative results, but Chung and colleagues47 and Lu and colleagues48 found a significant correlation between the Val66Met SNP and patients’ IQ. The Met allele was associated with lower IQ in patients with SSD in both studies, although in the case of Chung and colleagues,47 this relationship lost significance after Bonferroni correction.
Studies exploring the BDNF gene and IQ in patients with schizophrenia
DTNBP1
Five studies targeted polymorphisms at DTNBP1 (Table 3), of which 3 had a case–control methodology and 2 had a cross-sectional design. Two studies (40%) found no association between different SNPs in DTNBP1 and patients’ IQ;52,54 the remaining studies (60%) confirmed such an association.51,53,55 Burdick and colleagues51 observed that the genotype of 6 SNPs in DTNBP1 (rs909706, rs1018381, rs2619522, rs760761, rs2619528 and rs1011313) was associated with intellectual decline in patients: carriers of the CTCTAC haplotype demonstrated a significantly greater decline in IQ than noncarriers. Zinkstok and colleagues53 found that carriers of the low-frequency allele in SNPs rs760761 (T) and rs2619522 (G) had lower IQs, and the common allele in rs2619538 (A) was related to higher IQ. Varela-Gomez and colleagues55 found that patients who were homozygotes for the risk genotype in rs2619539 (GG) and in rs3213207 (AA) had lower IQs.
Studies exploring the DTNBP1 gene and IQ in patients with schizophrenia
TCF4
Three studies assessed the relationship between IQ in patients with SSD and polymorphisms in TCF4 (Table 4). Two of them (66.66 %) were cross-sectional and did not find a significant association.56,58 In contrast, a study57 with a case–control design observed that T-carrier patients at rs2958182 had higher IQs than A-carrier patients.
Studies exploring the TCF4 gene and IQ in patients with schizophrenia
Other candidate genes
Twenty-nine studies explored candidate genes other than those mentioned above; 26 had a case–control design, 2 were cross-sectional and 1 was longitudinal. Of these, 16 studies (55.17%) reported significant associations between the investigated polymorphisms and patients’ IQs in different directions (Table 5). In most cases, the minor frequent allele was associated with lower IQ in patients, including SNPs at ANK3,61 CSMD1,65 FOLH1,67 GRM7,68 GRM5,69 MHC,73 MIR137,74 NOS1,75 NRG1,77 NRGN78 and OXTR.82 For NRGN,78 an association with IQ was found only for the diplotype rs12807809–rs12278912, where the risk allele combination TG/TG was related to lower IQ. In contrast, the minor frequent allele in some SNPs at NRN1,79,80 TH85 and ZNF804A86,87 showed a protective effect: patients who were carriers had higher IQs. Two studies specified that the relationship between genetic variants and IQ was conditioned by the type of IQ. Donohoe and colleagues75 found that carriers of the risk genotype in rs6490121 (GG) at NOS1 had lower verbal IQs (both patients and controls), but full IQs were not significantly different. As well, some rare variants at OXTR demonstrated a specific link with low nonverbal IQ.82 Thirteen studies obtained nonsignificant results.59,60,62–64,66,70–72,76,81,83,84
Studies exploring different candidate genes and IQ in patients with schizophrenia
Genome-wide association studies
We included 6 GWASs in the review, of which 2 were cross-sectional and 4 had a case–control design (Table 6). Two studies explored SNPs and 4 searched CNVs. One of the SNP studies (50%) found no significant associations with patient IQ,88 but the other study89 reported 2 SNPs that were associated with patient IQ. The risk alleles were related to lower global IQ for the 2 SNPs located at genes CHD7 and GATAD2A;89 specifically, the risk allele in rs6984242 (CHD7) was the most strongly associated with low verbal IQ.
Overview of genome-wide association studies related to IQ and schizophrenia
From the remaining GWASs exploring CNVs, all reported significant associations were with low IQ. Martin and colleagues91 found that patients with large (> 500 kb) and rare (< 1 % frequency) deletions had lower IQs than those without such polymorphisms. Similarly, Lowther and colleagues35 observed that patients with pathogenic CNVs showed lower IQs, in contrast to patients with average IQs, who had the lowest yield of pathogenic CNVs. Derks and colleagues90 identified 14 CNVs, mostly at chromosome 15q11.2 (related to intellectual disability), after studying patients who had schizophrenia and IQ scores below 70. Finally, Hubbard and colleagues92 found that patients carrying CNVs related to SSD showed significantly lower IQs than noncarriers.
Discussion
This was, to our knowledge, the first systematic review to analyze current evidence related to the genetic association between SSD and IQ. We summarized 55 studies, some of which found that variability in several genes was significantly related to SSD and IQ in different directions. At the trait level, the results were consistent in showing lower estimated IQ in patients with SSD compared to healthy controls, corresponding to findings from other reviews and meta-analyses.93–95 Taken together, these findings show a possible common biological correlate between the 2 traits, indicating that IQ is a strong candidate endophenotype for SSD. Our review synthesizes the current data emerging from a line of research that has yielded heterogeneous results from diverse study strategies. For this reason, we believe that the present work provides relevant information that can lead researchers to select lines of investigation with the most promising evidence. Our results point to the study of novel genes such as NRN1, ZNF804A, CHD7 and GATAD2A, instead of others that have been more traditionally studied (COMT, BDNF and DTNBP1) but have not been supported by results from GWASs.
COMT
The most frequently studied polymorphism was Val158Met (rs4680), located at COMT. This gene encodes the enzyme catechol-O-methyltransferase (COMT), which degrades catecholamines, including dopamine, and helping to maintain appropriate levels of this neurotransmitter, particularly in the prefrontal cortex.96 The Val158Met polymorphism leads to a substitution of valine with methionine, in which the Val allele results in increased enzymatic activity of COMT, which in turn reduces dopamine concentrations.97 Some studies described here established that patients with the Val genotype showed lower IQs than Met carriers,43–45 but others did not replicate this finding.39–42 The possible IQ deficit in Val carriers might be explained by a dopamine reduction in the prefrontal cortex associated with this variant, which, in turn, is linked to worse performance in terms of executive function and working memory.98 Moreover, a study on intellectual disability suggested that Val158Met might contribute to intelligence by affecting white matter architecture in the prefrontal lobe and hippocampal formation.99
This discrepancy in findings could be the result of differences when controlling for mediating variables such as medication and type of intelligence. Rebollo-Mesa and colleagues43 noted that Val carriers had lower verbal IQs, but showed no differences in performance IQ, and this effect was exclusive for patients who were taking high doses of antipsychotics. This finding indicates that the Val158Met polymorphism could modulate the effects of antipsychotics on verbal IQ, in which carriers of the risk allele would be more susceptible to the deterioration of verbal skills as a result of medication.100 This corresponds with the findings of Schacht,101 who stated that this same polymorphism was key in identifying patients who were most likely to respond adequately to dopaminergic drugs. For clinical practice, these outcomes suggest that there is a subgroup of SSD patients with higher genetic risk of cognitive decline who need long-term follow-up.
Further research into the Val158Met polymorphism must explore possible interactions with other SNPs, such as rs1801133 at MTHFR, because these interactions may influence the expression of COMT.45 In fact, it has been demonstrated that polymorphisms in these 2 genes may be associated with IQ, especially because the MTHFR-T allele decreases the beneficial role of the COMT-Met allele in patients with schizophrenia.101
BDNF
All included studies on the BDNF gene analyzed the Val66Met polymorphism, probably because of its demonstrated link to schizophrenia102 and cognitive processes.103 This gene encodes brain-derived neurotrophic factor (BDNF), a neurotrophin that has a relevant role in neurodevelopment, synapse regulation and synaptic plasticity.104 Variations in this gene may lead to alterations in the BDNF protein that could cause impaired brain development and synapse and neuroplasticity failures, which have been associated with schizophrenia.104,105 The results found in this review related to the Val66Met polymorphism were controversial. Chung and colleagues47 and Lu and colleagues48 established that patients with SSD who were Met carriers had lower IQs than those who were Val carriers. This finding corresponded with previous research indicating that the BDNF-Met allele was associated with lower IQ in healthy women.106
The Val66Met polymorphism is believed to contribute to the etiology of SSD by affecting brain morphology as a result of lower levels of BDNF: Met allele carriers show reductions in the dorsolateral prefrontal cortex, caudate nucleus and frontal grey matter volume.107–109 The decreased secretion of BDNF caused by Val–Met substitution may also alter synaptic plasticity and neurodevelopment,105 which could influence cognition by disrupting the learning process.104 However, 3 other studies included in this review found no association between the Val66Met polymorphism and IQ in people with SSD.46,49,50 Once again, the different findings might depend on genetic variability in BDNF between populations, because a significant association was found only in individuals from South Korea and China. This finding was in line with previous literature reporting a higher frequency of the Met allele in the Asian population;110 future studies must explore the potential differential role of the Val66Met polymorphism in patients with SSD from different genetic backgrounds. Moreover, the association of Val66Met with IQ was established only for full IQ and verbal IQ estimated by the WAIS47,48 but not with premorbid IQ assessed using the NAART questionnaire.49 For this reason, Val66Met could be exclusively related to some types of IQ.
DTNBP1
Some associations were reported between SNPs at DTNBP1 and IQ in patients with SSD. This gene encodes the dysbindin-1 protein, which plays a relevant role in neurotransmission and neurodevelopment.111 Dysbindin-1 has been found in presynaptic and postsynaptic locations in several brain areas of interest in schizophrenia, including the hippocampus, the prefrontal cortex and the midbrain.112,113 Because dysbindin-1 interacts with different proteins involved in the release of neurotransmitters, its alteration could affect synaptic homeostasis.83 In all studies with significant results, the risk allele was related to lower IQ, including the C allele in rs2619539,55 the T allele in rs760761,53 the G allele in rs261952253 and the A allele in rs3213207.55 Furthermore, a risk haplotype that included 2 of the above SNPs (rs2619522 and rs760761) was associated with a greater decline in IQ in patients with SSD.51 The mechanism responsible for this association is unknown, but these polymorphisms might influence intelligence by reducing DTNBP1 expression in the prefrontal cortex, hippocampus and midbrain,111,112 thus affecting the glutamatergic system.113
However, other studies did not find this association,52,54 possibly because of the great variety of SNPs analyzed. In addition, although most studies considered patients in the early stages of SSD, Hashimoto and colleagues54 analyzed a sample with chronic SSD, which may have affected the results: changes in gene expression have been described depending on the clinical stage of the disorder.114 Similarly, the discrepancies found may have been a consequence of genetic variance in DTNBP1 between populations,113,115 because the only study with Asian participants had negative results.
TCF4
Few studies have explored the effect of genetic variability in TCF4 on IQ and SSD. TCF4 encodes transcription factor 4, a basic helix–loop–helix transcription factor. TCF4 is widely expressed in the early human embryo and might be relevant for nervous system development116 because of its role in neural proliferation and differentiation.117 Disruptions in this gene are associated with neurodevelopmental disorders that occur with intellectual disability, such as Pitt–Hopkins syndrome.116 However, this review found insufficient evidence to confirm a relationship between TCF4 variability and IQ in patients with SSD. A single study obtained significant findings,57 in which carriers of the minor allele in rs2958182 (A) had lower IQs than noncarriers. Lennertz and colleagues56 found that patients with SSD carrying the risk allele (C) of rs9960767 had worse memory impairment, and Albanna and colleagues58 observed that the risk allele was related to deficits in the reasoning cognitive domain. Therefore, TCF4 polymorphisms could be linked to specific cognitive domains rather than general cognition, but further studies are needed to understand their role in the pathophysiology of SSD.
Other candidate genes
Among the results of other candidate genes, NRN1 and ZNF804A showed the strongest links with IQ in patients with SSD. The NRN1 gene encodes a protein from the neuritin family and is expressed in both embryonic development118 and the adult brain.119 It plays a role in neuronal differentiation, synapse formation and maturation, and synaptic plasticity.120,121 Because of these functions, it is believed that variations of this gene could confer risk for the disorder and for neurocognitive alterations.80 Two different studies observed that a haplotype (rs1475157 and rs9405890) at NRN1 was related to IQ in patients with SSD.79,80 Chandler and colleagues79 reported that the SNPs rs1475157 and rs9405890 had a selective influence on fluid intelligence, because carriers of the GA haplotype had lower fluid intelligence scores on both premorbid and current IQ tests. As well, variability in rs1475157 and rs9405890 was related to age at onset, which in turn modulated the long-term cognitive course of patients with SSD.122 Although these findings must be replicated, research into NRN1 variations could help to identify a subgroup of patients at increased risk of cognitive impairment.
Two other studies observed that the risk allele (A) in rs1344706, located at ZNF804A, was associated with high IQ in patients with schizophrenia.86,87 Walters and colleagues86 clarified that although patients carrying this allele had higher IQs and fewer cognitive deficits than noncarriers, they still showed cognitive impairments compared to healthy participants. This polymorphism might be implicated specifically in SSD because it has been more frequently identified in patients with the disorder,8,123,124 and it has been related to higher schizotypy scores in healthy individuals.125 Overall, these findings agree on the value of the rs1344706 polymorphism as a potential genetic marker for risk of psychosis. The specific function of ZNF804A is still unknown, but it encodes a zinc finger binding protein, a type of protein that participates in diverse roles, such as binding to DNA, transcriptional regulation and DNA–protein interactions.126,127 This gene is expressed in the fetal and adult human brain, including the medial temporal lobe, the dorsolateral prefrontal cortex, the hippocampus and the amygdala.128–130 A plausible hypothesis is that the risk allele in rs1344706 may be related to SSD by affecting the expression of ZNF804A and other genes relevant for neurodevelopment,128–130 and by disturbing brain connectivity.131
Genome-wide association studies
In this review, COMT, BDNF and DTNBP1 — the most frequent candidates explored in relation to IQ in patients with SSD — were not associated with such phenotypes using a GWAS strategy. There may be different reasons for the inconsistency of these results. First, because the 2 GWASs exploring SNPs included in the present review assessed patients with SSD,88,89 clinical heterogeneity could have diminished the statistical power to detect the effect of such genes.132 Moreover, their sample sizes were small, with limited statistical power, which could have interfered in the identification of genetic variants related to the traits of interest. Another plausible explanation is that the aforementioned genes (COMT, BDNF and DTNBP1) do not confer risk for SSD, which would be consistent with the larger GWASs in schizophrenia to date.8,9,133 Instead, other less explored common variations may better explain the genetic correlation between SSD and IQ reported by different GWAS meta-analyses with samples above 100 000 individuals. The studies by Hagenaars and colleagues,28 Savage and colleagues29 and the Brain Consortium134 have described substantial evidence of pleiotropy between schizophrenia and cognition, in which lower IQ134 and slower reaction28 time were genetically correlated with the disorder. Regarding intelligence, Smeland and colleagues135 identified 75 distinct genomic loci that may underlie the genetics overlapping with schizophrenia, and gene set analysis suggests that these loci are implicated in neurodevelopment, synaptic integrity and neurotransmission. Future studies are needed that focus on these candidate loci to delve into the biological processes that might lead to the clinical and cognitive characteristics of patients with SSD. Several authors have suggested that GWASs with large samples could improve phenotypic characterization by establishing stricter inclusion criteria.132,136,137 Furthermore, candidate gene studies can provide interesting insights by including homogeneous samples with similar symptoms and clinical manifestations among participants, focusing on unravelling the neurobiological basis of behaviour.
Otherwise, recent GWASs have also highlighted the involvement of epigenetic mechanisms involved in SSD and IQ. Whitton and colleagues89 explored a list of genes that are chromatin modulators of gene expression and candidate genes for schizophrenia risk. They found that 2 SNPs — 1 in CHD7 and 1 in GATAD2A — were related to lower IQ in patients with SSD. The strongest association was between rs6984242 (in CHD7) and verbal IQ, in which carriers of the risk allele (G) showed lower IQs. CHD7 encodes chromodomain helicase DNA binding protein 7 (CHD7), which participates in the organization of chromatin, making it relevant for the regulation of gene transcription, DNA repair, replication and recombination.138 GATAD2A encodes the protein GATA zinc finger domain containing 2A, a subunit of the nucleosome remodelling and histone deacetylation complex, and it represses gene expression.139,140 These findings indicate an interesting line of research that explores the possible involvement of genes with epigenetic regulation functions in the risk for SSD and cognitive dysfunction.
The GWASs on CNVs were consistent in showing a relationship between CNV burden and low IQ in patients with SSD.35,90,92,141 However, this finding should be interpreted with caution, because in a recent meta-analysis including 10 studies on patients with SSD, their unaffected relatives and unrelated controls found no evidence of an association between CNV burden and overall IQ.142 Instead, the authors observed that CNVs had greater effects on specific cognitive abilities such as memory and perceptual reasoning. The potential influence of CNVs on intellectual deficits in SSD should be confirmed in future studies with larger samples that estimate different types of IQ (e.g., verbal IQ or performance IQ).
Future directions in this line of research include analyzing the various cognitive functions within the IQ construct to investigate how genetic components affect each process separately. Future research should also improve the diversity of ancestry in the samples analyzed. Although some recent work is investigating the genetic architecture of schizophrenia in Latin American and East Asian populations,143,144 representation of other groups, including Africans, is still lacking.
Limitations
Despite the genetic associations between IQ and SSD described above, approximately 45% (n = 25) of the included studies reported an absence of such a relationship. These negative results could reflect a true lack of association but could also have been because of heterogeneity in the samples when including SSD patients with diverse diagnoses and characteristics. The effect of medication might have been another confounding factor, because not all primary studies controlled for it. The instruments used to measure IQ and the type of intelligence estimated could also have been a cause of heterogeneity. Even when the same instrument was administered (in most cases, the Wechsler Adult Intelligence Scale), some authors used the full scale and others exclusively used subscales of verbal intelligence or performance intelligence. In addition, the use of different measures makes it difficult to analyze the different cognitive functions that encompass the construct of intelligence. Furthermore, most studies excluded patients with IQs of less than 70, and because intellectual disability is related to CNV and rare mutations,35 relevant genetic information could have been lost. Other major limitations of some primary studies were sample size, the lack of a control group and the majority inclusion of those of Caucasian origin, leaving aside other population groups, affecting the generalizability of results to other underrepresented populations.
Regarding the systematic review itself, its main limitation was that we did not screen grey literature, so possible unpublished negative results may not have been included. Furthermore, we found few GWASs that could replicate the findings of candidate genes with larger samples and less probability of biases. This could have been because of our strict eligibility criteria, including the selection of original studies that addressed SSD diagnosis and IQ estimation simultaneously. Therefore, large and relevant GWASs such as the review by Smeland and colleagues,135 which combined data from separate samples (schizophrenia, bipolar disorder and general population), were excluded from our results. Our findings might have been improved by a broader search strategy in terms of eligibility criteria. Likewise, the number of studies we found for each candidate gene was limited in most cases, affecting the generalizability of the results and highlighting the need for further genetic studies that replicate previous findings. Nevertheless, the present review offered some insight into the genetic basis for SSD and IQ using a systematic methodology. In addition, because we did not establish a time limitation in our bibliographic search, we analyzed all existing evidence on this topic. Similarly, the development of a protocol and its registration before the literature search contributed to the prevention of bias in the selection of studies.
Conclusion
The association between genetic variants and IQ in patients with SSD reported in the present systematic review highlights previous findings on the polygenic nature of both intelligence and SSD. However, current evidence related to specific genes associated with IQ and SSD is inconclusive. Genes traditionally studied, such as COMT, BDNF and DTNBP1, have not been confirmed using a GWAS approach. Instead, novel genes have been targeted to understand the molecular basis underlying the IQ deficit in SSD, including NRN1, ZNF804A, CHD7 and GATAD2A. Overall, our results support a neurodevelopmental hypothesis for SSD, because there is evidence of genetic risk factors that predispose individuals to both low IQ and risk for psychosis. In addition, results for IQ deficits in patients with SSD compared to healthy individuals, and for genetic variants associated with IQ and SSD, suggest that IQ is a valid endophenotype for SSD. Therefore, susceptibility to cognitive deficits might be present from brain development and would not be only a consequence of the disease. Therefore, IQ estimation might help to detect a subgroup of individuals at risk for psychosis.
Footnotes
Competing interests: None declared.
Contributors: N. Murillo-García, E. Setién-Suero and R. Ayesa-Arriola designed the study. N. Murillo-García and S. Barrio-Martínez acquired the data, which N. Murillo-García, J. Soler, S. Papiol and M. Fatjó-Vilas analyzed. N. Murillo-García and S. Barrio-Martínez wrote the article, which N. Murillo-García, E. Setién-Suero, J. Soler, S. Papiol, M. Fatjó-Vilas and R. Ayesa-Arriola reviewed. All authors approved the final version to be published, agreed to be accountable for all aspects of the work and can certify that no other individuals not listed as authors have made substantial contributions to the paper.
Registry: International Prospective Registry of Systematic Reviews (PROSPERO) CRD42020218842.
Funding: This work was supported by a “Miguel Servet” contracts (R. Ayesa-Arriola and M. Fatjó-Vilas) from the Carlos III Health Institute (CP18/00003 and CP20/00072, respectively); a “Juan de la Cierva-Formación” contract (E. Setién-Suero) from the Spanish Ministry of Science and Innovation (FJC2019-042390-I/AEI/10.13039/501100011033); and a predoctoral contract (N. Murillo-García) from the Valdecilla Biomedical Research Institute and the University of Cantabria (BOC49, 19 REF. IDI-13).
Data availability: The data supporting this systematic review are available by request from the corresponding author RAA.
- Received February 14, 2022.
- Revision received June 27, 2022.
- Revision received August 19, 2022.
- Accepted September 6, 2022.
This is an Open Access article distributed in accordance with the terms of the Creative Commons Attribution (CC BY-NC-ND 4.0) licence, which permits use, distribution and reproduction in any medium, provided that the original publication is properly cited, the use is noncommercial (i.e., research or educational use), and no modifications or adaptations are made. See: https://creativecommons.org/licenses/by-nc-nd/4.0/