Abstract
As the tissue macrophages of the CNS, microglia are critically involved in diseases of the CNS. However, it remains unknown what controls their maturation and activation under homeostatic conditions. We observed substantial contributions of the host microbiota to microglia homeostasis, as germ-free (GF) mice displayed global defects in microglia with altered cell proportions and an immature phenotype, leading to impaired innate immune responses. Temporal eradication of host microbiota severely changed microglia properties. Limited microbiota complexity also resulted in defective microglia. In contrast, recolonization with a complex microbiota partially restored microglia features. We determined that short-chain fatty acids (SCFA), microbiota-derived bacterial fermentation products, regulated microglia homeostasis. Accordingly, mice deficient for the SCFA receptor FFAR2 mirrored microglia defects found under GF conditions. These findings suggest that host bacteria vitally regulate microglia maturation and function, whereas microglia impairment can be rectified to some extent by complex microbiota.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Ransohoff, R.M. & Perry, V.H. Microglial physiology: unique stimuli, specialized responses. Annu. Rev. Immunol. 27, 119–145 (2009).
Kettenmann, H., Hanisch, U.K., Noda, M. & Verkhratsky, A. Physiology of microglia. Physiol. Rev. 91, 461–553 (2011).
Prinz, M. & Priller, J. Microglia and brain macrophages in the molecular age: from origin to neuropsychiatric disease. Nat. Rev. Neurosci. 15, 300–312 (2014).
Schafer, D.P. & Stevens, B. Phagocytic glial cells: sculpting synaptic circuits in the developing nervous system. Curr. Opin. Neurobiol. 23, 1034–1040 (2013).
Naj, A.C. et al. Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer′s disease. Nat. Genet. 43, 436–441 (2011).
Hollingworth, P. et al. Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer's disease. Nat. Genet. 43, 429–435 (2011).
Guerreiro, R.J. et al. Using exome sequencing to reveal mutations in TREM2 presenting as a frontotemporal dementia-like syndrome without bone involvement. JAMA Neurol. 70, 78–84 (2013).
Rademakers, R. et al. Mutations in the colony stimulating factor 1 receptor (CSF1R) gene cause hereditary diffuse leukoencephalopathy with spheroids. Nat. Genet. 44, 200–205 (2012).
Zhan, Y. et al. Deficient neuron-microglia signaling results in impaired functional brain connectivity and social behavior. Nat. Neurosci. 17, 400–406 (2014).
Prinz, M. & Mildner, A. Microglia in the CNS: immigrants from another world. Glia 59, 177–187 (2011).
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010).
Schulz, C. et al. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science 336, 86–90 (2012).
Kierdorf, K. et al. Microglia emerge from erythromyeloid precursors via Pu.1- and Irf8-dependent pathways. Nat. Neurosci. 16, 273–280 (2013).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Goldmann, T. et al. A new type of microglia gene targeting shows TAK1 to be pivotal in CNS autoimmune inflammation. Nat. Neurosci. 16, 1618–1626 (2013).
Prinz, M., Tay, T.L., Wolf, Y. & Jung, S. Microglia: unique and common features with other tissue macrophages. Acta Neuropathol. 128, 319–331 (2014).
Grenham, S., Clarke, G., Cryan, J.F. & Dinan, T.G. Brain-gut-microbe communication in health and disease. Front Physiol 2, 94 (2011).
Gaspar, P., Cases, O. & Maroteaux, L. The developmental role of serotonin: news from mouse molecular genetics. Nat. Rev. Neurosci. 4, 1002–1012 (2003).
Diaz Heijtz, R. et al. Normal gut microbiota modulates brain development and behavior. Proc. Natl. Acad. Sci. USA 108, 3047–3052 (2011).
Bercik, P. et al. The intestinal microbiota affect central levels of brain-derived neurotropic factor and behavior in mice. Gastroenterology 141, 599–609, 609 (2011).
Dorrestein, P.C., Mazmanian, S.K. & Knight, R. Finding the missing links among metabolites, microbes, and the host. Immunity 40, 824–832 (2014).
Kamada, N., Seo, S.U., Chen, G.Y. & Nunez, G. Role of the gut microbiota in immunity and inflammatory disease. Nat. Rev. Immunol. 13, 321–335 (2013).
Round, J.L. & Mazmanian, S.K. The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9, 313–323 (2009).
Markle, J.G. et al. gammadelta T cells are essential effectors of type 1 diabetes in the nonobese diabetic mouse model. J. Immunol. 190, 5392–5401 (2013).
Wu, H.J. et al. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity 32, 815–827 (2010).
Berer, K. et al. Commensal microbiota and myelin autoantigen cooperate to trigger autoimmune demyelination. Nature 479, 538–541 (2011).
Reikvam, D.H. et al. Depletion of murine intestinal microbiota: effects on gut mucosa and epithelial gene expression. PLoS ONE 6, e17996 (2011).
Kierdorf, K. & Prinz, M. Factors regulating microglia activation. Front. Cell. Neurosci. 7, 44 (2013).
Sofer, A., Lei, K., Johannessen, C.M. & Ellisen, L.W. Regulation of mTOR and cell growth in response to energy stress by REDD1. Mol. Cell. Biol. 25, 5834–5845 (2005).
Hickman, S.E. et al. The microglial sensome revealed by direct RNA sequencing. Nat. Neurosci. 16, 1896–1905 (2013).
Chiu, I.M. et al. A neurodegeneration-specific gene-expression signature of acutely isolated microglia from an amyotrophic lateral sclerosis mouse model. Cell Reports 4, 385–401 (2013).
Butovsky, O. et al. Identification of a unique TGF-beta–dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Greter, M. et al. Stroma-derived interleukin-34 controls the development and maintenance of langerhans cells and the maintenance of microglia. Immunity 37, 1050–1060 (2012).
Khosravi, A. et al. Gut microbiota promote hematopoiesis to control bacterial infection. Cell Host Microbe 15, 374–381 (2014).
Prinz, M. et al. Microglial activation by components of gram-positive and -negative bacteria: distinct and common routes to the induction of ion channels and cytokines. J. Neuropathol. Exp. Neurol. 58, 1078–1089 (1999).
Cunningham, C., Wilcockson, D.C., Campion, S., Lunnon, K. & Perry, V.H. Central and systemic endotoxin challenges exacerbate the local inflammatory response and increase neuronal death during chronic neurodegeneration. J. Neurosci. 25, 9275–9284 (2005).
Stecher, B. et al. Like will to like: abundances of closely related species can predict susceptibility to intestinal colonization by pathogenic and commensal bacteria. PLoS Pathog. 6, e1000711 (2010).
Schaedler, R.W. & Dubos, R.J. The fecal flora of various strains of mice. Its bearing on their susceptibility to endotoxin. J. Exp. Med. 115, 1149–1160 (1962).
Bindels, L.B., Dewulf, E.M. & Delzenne, N.M. GPR43/FFA2: physiopathological relevance and therapeutic prospects. Trends Pharmacol. Sci. 34, 226–232 (2013).
Smith, P.M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
Chu, H. & Mazmanian, S.K. Innate immune recognition of the microbiota promotes host-microbial symbiosis. Nat. Immunol. 14, 668–675 (2013).
Clarke, G. et al. The microbiome-gut-brain axis during early life regulates the hippocampal serotonergic system in a sex-dependent manner. Mol. Psychiatry 18, 666–673 (2013).
Fiedler, K., Kokai, E., Bresch, S. & Brunner, C. MyD88 is involved in myeloid as well as lymphoid hematopoiesis independent of the presence of a pathogen. Am. J. Blood Res. 3, 124–140 (2013).
Kawai, T. & Akira, S. The roles of TLRs, RLRs and NLRs in pathogen recognition. Int. Immunol. 21, 317–337 (2009).
Deshmukh, H.S. et al. The microbiota regulates neutrophil homeostasis and host resistance to Escherichia coli K1 sepsis in neonatal mice. Nat. Med. 20, 524–530 (2014).
Borre, Y.E. et al. Microbiota and neurodevelopmental windows: implications for brain disorders. Trends Mol. Med. 20, 509–518 (2014).
Braniste, V. et al. The gut microbiota influences blood-brain barrier permeability in mice. Sci. Transl. Med. 6, 263ra158 (2014).
Schéle, E. et al. The gut microbiota reduces leptin sensitivity and the expression of the obesity-suppressing neuropeptides proglucagon (Gcg) and brain-derived neurotrophic factor (Bdnf) in the central nervous system. Endocrinology 154, 3643–3651 (2013).
Perez-Burgos, A. et al. Psychoactive bacteria Lactobacillus rhamnosus (JB-1) elicits rapid frequency facilitation in vagal afferents. Am. J. Physiol. Gastrointest. Liver Physiol. 304, G211–G220 (2013).
Giovanoli, S. et al. Stress in puberty unmasks latent neuropathological consequences of prenatal immune activation in mice. Science 339, 1095–1099 (2013).
Conrad, M.L. et al. Maternal TLR signaling is required for prenatal asthma protection by the nonpathogenic microbe Acinetobacter lwoffii F78. J. Exp. Med. 206, 2869–2877 (2009).
Dann, A. et al. Cytosolic RIG-I-like helicases act as negative regulators of sterile inflammation in the CNS. Nat. Neurosci. 15, 98–106 (2012).
Mildner, A. et al. Microglia in the adult brain arise from Ly-6C(hi)CCR2(+) monocytes only under defined host conditions. Nat. Neurosci. 10, 1544–1553 (2007).
Raasch, J. et al. I{kappa}B kinase 2 determines oligodendrocyte loss by non-cell-autonomous activation of NF-{kappa}B in the central nervous system. Brain 134, 1184–1198 (2011).
Jaitin, D.A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Cahenzli, J., Koller, Y., Wyss, M., Geuking, M.B. & McCoy, K.D. Intestinal microbial diversity during early-life colonization shapes long-term IgE levels. Cell Host Microbe 14, 559–570 (2013).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Eden, E., Lipson, D., Yogev, S. & Yakhini, Z. Discovering motifs in ranked lists of DNA sequences. PLoS Comput. Biol. 3, e39 (2007).
Herz, J. et al. Acid sphingomyelinase is a key regulator of cytotoxic granule secretion by primary T lymphocytes. Nat. Immunol. 10, 761–768 (2009).
Acknowledgements
This work is dedicated to our former friend Uwe-Karsten Hanisch, past Professor for Neuroscience at the Paul-Flechsig-Institute for Brain Research, Leipzig, who devoted his whole life to the exploration of microglia function. We thank M. Oberle, M. Ditter and T. el Gaz for excellent technical assistance. We are grateful to J. Bodinek-Wersing and G. Rappl for cell sorting and to A. Spies for providing cDNA from cell cultures. M.P. was supported by the BMBF-funded competence network of multiple sclerosis (KKNMS), the Sobek Foundation, the DFG (SFB 992, FOR1336, PR 577/8-1), the Fritz-Thyssen Foundation and the Gemeinnützige Hertie Foundation (GHST). B.S. was supported by the SPP1656 “Intestinal Microbiota” (STE 1971/4-1).
Author information
Authors and Affiliations
Contributions
D.E., A.L.H.d.A., P.W., D.J., K.J., T.M., B.S., O.U., I.A., H.K.-S., E.D. and K.D.M. conducted the experiments. I.A. and O.S. analyzed the RNA sequencing and microarray data, respectively. T.B., B.S., A.D., P.S., V.S., E.C. and W.S.G. provided mice and designed experiments. M.P. supervised the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
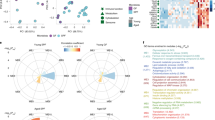
Supplementary Figure 1 Gene profile from microglia in GF mice.
mRNA expression values (number of reads) of genes from microglia in GF animals (white bars) or SPF mice (black bars). (a) Prototypical microglia genes are presented and compared to the non-microglia genes such as Gfap, Mog, Tubb3 and Rbfox3. (b) Epigenetic signature of microglia was examined by investigating genes (number of reads) related to histone acetylation, deacetylation, methylation and demethylation, respectively. Bars represent means ± s.e.m. with seven samples in each group. Significant differences were determined by an unpaired t test and marked with asterisks (*P < 0.05, **P < 0.01, ***P < 0.001). P values: Csf1r: 0.0001, Sfpi1: 0.0075, Hexb: 0.0060. Myst3: 0.0121, Hdac1: 0.0191, Sirt2: 0.0113, Sirt7: 0.0441, Mll3: 0.0489, Mll5: 0.0105, Setd1a: 0.0360, Setd7: 0.0134, Kdm4a: 0.0476, Kdm6b: 0.0005, Jarid2: <0.0001.
Supplementary Figure 2 Absence of microbes does not alter the amount of neurons, astrocytes and oligodendrocytes.
CNS histology of the neuroectodermal compartment of 6-8 weeks old GF or control SPF mice. NeuN labels mature neurons, Nogo-A oligodendrocytes and glial fibrillary acidic protein (GFAP) astrocytes. Representative pictures are shown. Scale bars: NeuN: 100 µm, Nogo-A and GFAP: 50 µm.
Supplementary Figure 3 Absence of microbes does not alter the amount of neurons, astrocytes and oligodendrocytes.
(a,b) Quantitative assessment of the neuroectodermal compartment of 6-8 weeks old GF or control SPF mice. Hippocampal NeuN+ cells were counted in the gyrus dentatus. Cerebellar NeuN+ cells were counted in the granular cell layer. Each symbol represents one mouse, with five samples per group. Means ± s.e.m. are indicated. No significant differences could be determined by using an unpaired t test. Representative pictures are shown. n.d. = not detectable. Data are representative of two independent experiments. Scale bars: H&E: 200 µm.
Supplementary Figure 4 Lymphocytic cells are indistinguishable in the brains of SPF and GF mice.
Immunohistochemistry for T cells (CD3) and B cells (B220) in different brain regions of SPF and GF mice, respectively. Insets depict spleen sections as positive controls. Scale bars: overviews: cortex, corpus callosum and cerebellum (CD3): 50 µm; olfactory bulb (CD3): 20 µm; hippocampus (CD3), cortex, corpus callosum, hippocampus, olfactory bulb and cerebellum (B220): 100 µm. Inset: 50 µm (CD3) and 100 µm (B220).
Supplementary Figure 5 Increased microglia number in cerebellar grey and white matter of GF mice.
(a) Quantification of Iba-1+ parenchymal microglia from either SPF (black symbols) or GF (white symbols) mice was performed on cerebellar brain slices. Each symbol represents one mouse. Three to four sections per mouse were examined. Means ± s.e.m. are shown. Significant differences were determined by an unpaired t test and marked with asterisks (*P < 0.05, ***P < 0.001). P values: granular cell layer: 0.0008, white matter: 0.0287. Representative pictures of nine examined mice per group are displayed.
(b) Imaris-based 3-D reconstruction of the cortical microglia networks under SPF (left) or GF (right) conditions. GF microglia have no distances to neighbouring microglia and show partial physical contacts to adjacent cells. Inset exhibits such an interaction. Scale bars: 20 µm (overview), 3 µm (inset).
(c) Quantitative RT-PCR for the indicated mRNAs. Data are expressed as ratio of mRNA expression versus endogenous Actb relative to SPF conditions and exhibited as mean ± s.e.m. Bars (black: SPF, white: GF) represent means ± s.e.m. with at least four samples in each group. Data are representative of two independent experiments. Significant differences were determined by an unpaired t test and marked with asterisks (*P < 0.05, **P < 0.01, ***P < 0.001). n.d. = not detectable. P values: Il-34: hippocampus: 0.0031. Csf1: cortex: 0.0071, cerebellum: 0.0009. Tgfß1: cortex: 0.0253, cerebellum: 0.031. Sfpi1: cortex: 0.0302, corpus callosum: 0.0290, cerebellum: 0.0345.
Supplementary Figure 6 Decreased inflammatory repertoire of microglia from GF mice upon intracerebral LPS challenge.
Gene ontology enrichment network on differentially expressed genes in microglia from SPF (a) or GF (b) mice on the basis of an Affymetrix DNA microarray analysis. Diagram depicts functional results of GO clustering through GORilla. Only very highly significantly overrepresented GO terms are included with P values ranging from P < 10-9 (yellow) to P < 10-24 (red). White framed terms depict not significantly overrepresented parent terms of the colored terms.
Supplementary Figure 7 Diminished inflammatory response of microglia from GF mice upon peripheral LPS challenge.
Quantitative RT-PCR of LPS-induced factors in microglia 6 hrs after intraperitoneal LPS challenge. Data are expressed as ratio of the mRNA expression compared to endogenous Actb relative to control and show mean ± s.e.m. At least three mice per group were analyzed. Data are representative of two independent experiments. Significant differences were determined by an unpaired t test and marked with asterisks (*P < 0.05, **P < 0.01, ***P < 0.001). ns: not significant. P values: PBS: Il1- b:0.3720, Il-6: 0.5625, Il12 b: 0.2442, Tnfα p=0.5955, Marco: 0.0642, Ccl2: 0.6608, Ccl7: 0.3316, Cxcl10: 0.0867, Csf1: 0.1194, cyclin D2: 0.1135, S100a4: 0.5357, S100a6: 0.4281, S100a8: 0.0642, S100a9: 0.4583, S100a10: 0.1771. LPS: Il-1 b: 0.0023, Il-6: 0.1329, Il-12 b: <0.0001, Tnfα: 0.0008, Marco: 0.0001, Ccl2: 0.0673, Ccl7: 0.2708, Cxcl10:.1522, Csf1: 0.1828, cyclin D2: 0.0344, S100a4: 0.0064, S100a6: 0.0341, S100a8: 0.0009, S100a9: 0.5782, S100a10: 0.0352.
Supplementary Figure 8 Determination of intestinal bacterial loads and complexity.
(a) qPCR analysis for eubacterial 16S rRNA could not detect significant amounts of bacteria in the caeca of GF and antibiotic-treated SPF animals. Representative data from fifteen SPF, five GF, six SPF (ABX), five ASF and five ASF (recol.) mice are depicted. Gene copy numbers were normalized to DNA content. Each symbol represents one mouse. Means ± s.e.m. are indicated. Significant differences were determined by an unpaired t test and marked with asterisks (*P < 0.05, **P < 0.01). P values: SPF vs. GF: 0.0151, SPF vs. SPF (ABX): 0.0046, SPF vs. ASF (recol.): 0.0280.
(b) Recolonized ASF mice (ASF recol.) harbour a greater microbial diversity as detected by qPCR using group specific primers. Besides Parabacteroides (ASF357), Lactobacillus (ASF 361) and Clostridium (ASF356), mouse intenstinal Bacteroides (mib) and Enterobacteriaceae groups are detectable in the caecum of SPF and re-colonised ASF animals using group-specific primers, respectively.
Supplementary Figure 9 Microglia maturation is independent from the TLRs2,3,4,7,9.
(a) Left: View of caecum from a SPF-TLRwt (control, left) and SPF-TLR2,3,4,7,9-deficient (SPF-TLRko) mouse (right). Ruler scale is shown. Representative pictures are shown. Right: Relative caecum weight of control SPF-TLRwt and SPF-TLRko mice. Symbols represent individual mice, with six animals per group. Means ± s.e.m. are shown. No statistically significant differences were found by an unpaired t test.
(b) Iba-1 immunohistochemistry in the cortex of SPF-TLRwt and SPF-TLRko animals (left) and quantification thereof (right). Every symbol represents one mouse. Six animals per group were analyzed and three to four sections per mouse were examined. Data are representative of two independent experiments. Means ± s.e.m. are indicated. No significant differences could be determined by an unpaired t test.
(c) Ddit4 mRNA measured by qRT-PCR in microglia from SPF-TLRwt (black bar) or SPF-TLRko (white bar) mice. Means ± s.e.m. with six samples in each group are displayed. No significant differences could be determined by an unpaired t test. Data are representative of two independent experiments.
(d) Three-dimensional reconstruction (Scale bars: 15µm) and Imaris-based automatic quantification of cell morphometry (e) of Iba-1+ microglia. Individual symbols represent one mouse per group with at least three measured cells per mouse. Six mice per group were analyzed. Means ± s.e.m. are indicated. No significant differences were detected by an unpaired t test.
(f) Flow cytometry of isolated microglia from control six SPF-TLRwt and six SPF-TLRko individuals. Percentages and mean fluorescence intensities (MFI) of the indicated surface molecules on microglia are shown. Each symbol represents one mouse. Means ± s.e.m. are indicated. Significant differences were evaluated by an unpaired t test and labelled with asterisks (*P < 0.05). P value: CD31 (MFI): 0.0419. Data are representative of two independent experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 (PDF 1810 kb)
Supplementary Table 1: Altered microglial gene profile and immaturity in germ-free animals.
Significantly down- (upper table) and up- (lower table) regulated genes subjected to functional GO cluster analysis. The significance of genes is indicated by the given P values. (XLS 32 kb)
Supplementary Table 2: Reduced gene induction in microglia from non-colonized GF animals.
Affymetrix Genome 430A 2.0 array-based gene induction of microglia from SPF mice compared to GF individuals. 78 genes significantly differentially expressed in GF vs. SPF microglia 6 hr after i.c. LPS treatment are shown. Only genes also significantly up- or downregulated by LPS treatment compared to PBS treated controls of the same housing conditions (GF and SPF respectively) are included to account for differences in basal gene regulation. (P < 0.01). (XLS 41 kb)
Rights and permissions
About this article
Cite this article
Erny, D., Hrabě de Angelis, A., Jaitin, D. et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat Neurosci 18, 965–977 (2015). https://doi.org/10.1038/nn.4030
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4030