Abstract
Background: Brain imaging is of limited diagnostic use in psychiatry owing to clinical heterogeneity and low sensitivity/specificity of between-group neuroimaging differences. Machine learning (ML) may better translate neuroimaging to the level of individual participants. Studying unaffected offspring of parents with bipolar disorders (BD) decreases clinical heterogeneity and thus increases sensitivity for detection of biomarkers. The present study used ML to identify individuals at genetic high risk (HR) for BD based on brain structure.
Methods: We studied unaffected and affected relatives of BD probands recruited from 2 sites (Halifax, Canada, and Prague, Czech Republic). Each participant was individually matched by age and sex to controls without personal or family history of psychiatric disorders. We applied support vector machines (SVM) and Gaussian process classifiers (GPC) to structural MRI.
Results: We included 45 unaffected and 36 affected relatives of BD probands matched by age and sex on an individual basis to healthy controls. The SVM of white matter distinguished unaffected HR from control participants (accuracy = 68.9%, p = 0.001), with similar accuracy for the GPC (65.6%, p = 0.002) or when analyzing data from each site separately. Differentiation of the more clinically heterogeneous affected familiar group from healthy controls was less accurate (accuracy = 59.7%, p = 0.05). Machine learning applied to grey matter did not distinguish either the unaffected HR or affected familial groups from controls. The regions that most contributed to between-group discrimination included white matter of the inferior/middle frontal gyrus, inferior/middle temporal gyrus and precuneus.
Limitations: Although we recruited 126 participants, ML benefits from even larger samples.
Conclusions: Machine learning applied to white but not grey matter distinguished unaffected participants at high and low genetic risk for BD based on regions previously implicated in the pathophysiology of BD.
Introduction
The diagnostic system in psychiatry continues to be based on behavioural symptoms rather than biomarkers. This complicates clinical work and research as it introduces marked heterogeneity. Neuroimaging has the unique ability to noninvasively investigate brain structure and function. Yet, the diagnostic promise of neuroimaging in psychiatry has not been fully realized. Brain imaging studies have shown replicated evidence for neuroanatomical changes in groups of participants with psychiatric disorders relative to controls. However, these statistical group differences have low specificity and sensitivity and thus are of limited diagnostic use on the level of individual participants.1–3
The problem of low sensitivity and specificity may be overcome by novel methods of neuroimaging analyses, such as machine learning (ML).1,2 Traditional methods of MRI data analysis focus on relatively large, localized and spatially segregated patterns of between-group differences.4 In contrast, the multivariate ML techniques target patterns of relatively minor alterations distributed throughout the whole brain,1 which may better characterize the abnormalities found in individuals with psychiatric disorders.5 These techniques bring neuroimaging analyses to the level of individual participants and potentially allow for their diagnostic use.
The use of neuroimaging for diagnostic purposes in psychiatry is further complicated by clinical heterogeneity.6,7 Not all neuroimaging findings in psychiatric patients are of diagnostic use. For example, brain changes in patients with bipolar disorders (BD) may represent biological markers of BD, but also the consequences of illness episodes,8,9 exposure to medications10–12 or comorbid conditions.13,14 The changes secondary to illness burden, medication exposure or comorbid conditions have limited diagnostic potential as they occur only later in the course of BD.
The biomarkers that could be used diagnostically (i.e., the brain changes that reflect the susceptibility for BD) are typically small.15–17 They may be underrepresented in convenience samples, which include patients with long chronic illness, polypharmacy and medical and psychiatric comorbid conditions.18 In such samples, neuroimaging may primarily detect the consequences of the illness, medication exposure or signatures of comorbid conditions, which may mask or override the diagnostically relevant biomarkers.8,12,16,18,19 Consequently, the identification of brain changes with diagnostic potential requires research designs that attempt to minimize or completely eliminate the secondary brain changes and thus increase sensitivity for detection of the primary alterations/biological risk factors. This can be achieved by studying the unaffected offspring of parents with BD, who are at high genetic risk for the illness but do not show any consequences of the illness — a so-called genetic high risk (HR) design. Any changes detected among unaffected, medication-naive individuals cannot be a consequence of illness burden, treatment or comorbid conditions. Such alterations, especially if replicated among the affected participants, likely reflect susceptibility for BD, which may be of diagnostic utility.17,20
Combining ML with an HR study is a particularly strong research design. By decreasing clinical heterogeneity, it increases the sensitivity for detection of biomarkers with diagnostic potential.17,20 Yet, to our knowledge, this approach has not previously been used in BD. To fill this knowledge gap, we tested the feasibility of applying ML to differentiate patients at risk for BD from healthy controls based on brain structure.
Methods
Study design
We recruited offspring from families of well-characterized adult BD probands in 2 centres: Halifax, NS, Canada, and Prague, Czech Republic (2-centre HR design). The details about the recruitment and samples have been published previously.18,21,22 We divided the participants based on the presence or absence of personal history of mood disorders. Including both affected and unaffected offspring is necessary to establish the presence of neurobiological changes in families and their association with the illness. The average genetic liability among unaffected offspring of BD probands decreases with age, as those with higher liability become affected. Therefore, it is important to include individuals around the typical age of onset, who remain at a substantial risk of future conversion to BD.23,24 Thus, the inclusion criterion for all groups in both centres was age between 15 and 30 years. Common exclusion criteria for both groups in both centres were personal history of any serious medical/neurologic disorders, substance abuse/dependence during the last 6 months and MRI exclusion criteria. In addition to these, controls from both centres were excluded if they had any personal or family history of DSM-IV Axis I psychiatric disorders.
Probands
Families were identified via adult probands with BD who had participated in 1 of the following: previous genetic studies for the Halifax sample or the Czech Bipolar Disorder Case Registry22 for the Prague sample. The probands completed the Schedule for Affective Disorders and Schizophrenia — Lifetime version (SADS-L)25 interview, which was conducted by board certified psychiatrists. Final DSM-IV diagnoses were derived using all available clinical materials in a blind consensus fashion by an independent panel of senior clinical researchers.
High-risk offspring
The offspring from the families described above were interviewed by child/adolescent or adult psychiatrists using the Schedule for Affective Disorders and Schizophrenia for School-Age Children KSADS-PL26 or SADS-L format, depending on their age. We divided the offspring into 2 groups: the HR unaffected group or the affected familial group.
The HR unaffected group consisted of 50 offspring with no lifetime history of psychiatric disorders. These individuals were at an increased risk for BD because they had 1 parent affected with a primary mood disorder. In general the risk for BD developing among offspring of parents with BD is about 10 times greater than the risk in the general population, and up to 50% of BD offspring may experience some form of psychiatric morbidity.27 In our sample of prospectively followed individuals, among whom participants for the present study were recruited, the age-adjusted prevalence of major mood disorders reached 53% by an average age of 20 years.28
The affected familial group consisted of 36 offspring who met the criteria for a lifetime Axis I diagnosis of mood disorders (i.e., a personal history of at least 1 episode of depression, hypomania or mania meeting full DSM-IV criteria). Unipolar depression among offspring of parents with BD is typically the first manifestation of BD,29 which was also the case in our sample of prospectively followed individuals, among whom participants for the present study were recruited.23,28
Control group (offspring of healthy parents)
We also recruited 49 healthy offspring from families without any personal or family history of psychiatric disorders. These individuals had similar characteristics to the experimental groups regarding age, sex and sociodemographic background. They were interviewed by a child/adolescent or adult psychiatrist according to a KSADS-PL or SADS-L format, depending on their age, and determined to be free of psychiatric illness. Negative psychiatric family history was evaluated by acquiring family history from the participants, and if possible, 1 of their parents. The controls were matched on an individual basis by age (within 1 year) and sex to the unaffected and affected offspring. Five of the unaffected HR offspring did not have a matching control, yielding a sample of 45 unaffected HR participants. All of the 36 affected participants had a matching control.
Prior to conducting the assessments, all interviewers underwent extensive training consisting of participation in interviews, interviews under supervision and blind consensus diagnostic reviews.
After providing a complete description of the study, written informed consent was obtained from every individual. The studies were approved by the research ethics boards of the IWK Health Centre and the Capital District Health Authority in Halifax, NS, and by The Prague Psychiatric Center Institutional Review Board.
MRI acquisition parameters
The participants were scanned at the 2 sites. We used the same scanner type and scanning parameters at both sites. Thus, all MRI acquisitions were performed with a 1.5 T General Electric Signa scanner and a standard single-channel head coil. After a localizer scan, a T1-weighted spoiled gradient recalled (SPGR) scan was acquired with the following parameters: flip angle 40°, echo time 5 ms, repetition time 25 ms, field of view 24 cm × 18 cm, matrix 256 × 160 pixels, number of excitations = 1, no interslice gap, 124 coronal, 1.5 mm thick slices.
Data preprocessing
Similar to other ML studies,30–32 the data were preprocessed with SPM8 software (Wellcome Department of Imaging Neuroscience Group; www.fil.ion.ucl.ac.uk/spm) and the VBM8 toolbox (http://dbm.neuro.uni-jena.de/vbm.html) using default parameters and following standard methods, as used previously by our group18 and others.33,34 Specifically, the images were bias-corrected, tissue classified and registered using linear (12-parameter affine) and nonlinear transformations (warping), within a unified model.35 The resulting images were visually inspected for quality by expert raters blinded to group assignment and guided by boxplots and covariance matrices provided by the VBM8 toolbox. There were no visually identifiable excessive motion artifacts in the data. Normalized and modulated grey matter and white matter segmented images were then smoothed with 8 mm isotropic Gaussian kernels and used as input into the classification algorithms. A mask was applied including only grey matter or only white matter voxels in common for all participants (voxels with grey or white matter probability value equal to zero for at least 1 participant were excluded from the respective analyses). This is the most common method of structural MRI data preprocessing for ML analyses.
Support vector machines and Gaussian process classifiers
The pattern classification analyses were performed using the PROBID toolbox (www.brainmap.co.uk/probid.htm).
The support vector machines (SVM) and Gaussian process classifiers (GPC) are 2 standard methods of ML pattern recognition, which have previously been applied to analyses of structural MRI in psychiatry.1,30 Technical descriptions of GPC and SVM inference have been presented elsewhere.31,36,37,38 Similar to other studies,30,32 we used the default parameters for the SVM and GPC analyses. This reduces methodological heterogeneity, ensures comparability between the studies and reduces the risk of overfitting.
The SVM classifier is trained by providing examples of the form < x,c > where x represents a spatial pattern (e.g., grey matter image) and c is the class label (e.g., c = +1 for unaffected HR participants and c = −1 for controls). During the training phase, the SVM finds the hyperplane or decision function that separates the examples in the input space according to the group label (e.g., HR participants v. low risk controls). We used a linear kernel SVM, which is less prone to overfitting than nonlinear SVMs. Linear kernel SVMs have a single parameter, C, that controls the trade-off between having zero training errors and allowing misclassifications. Similar to most other studies30,32,39 this was fixed at C = 1, which is the default value. The SVM performance for whole brain classification does not change for a large range of C values and degrades only with very small C values.40 Modifying the C threshold is suggested only when the dimensionality of the data is smaller than the number of examples (e.g., classification based on small regions of interest), which was not the case in our study. For the SVM, the optimal hyperplane is described by a weight vector and an offset.
In contrast to the categorical SVM method, the GPC classifier determines a predictive distribution that best distinguishes cases from controls. Once the decision function is determined from the training data, it can be used to predict the group membership of a new test example. The results of GPC are predictive probabilities scaled between zero and 1 that precisely quantify the predictive uncertainty of the classifier for the test case.
Cross-validation
We trained the GPC and SVM classifiers independently in each site and for the combined data set. The performance of each classifier was then validated with the commonly used “leave 2 out” cross-validation approach, which provides a relatively unbiased estimate of the true performance.39 To allow for this, we matched participants on an individual basis by age and sex to controls. In each trial, observations from all but 1 participant from each group were used to train the classifier. Subsequently, the class assignment of the test participants was calculated during the test phase. This procedure was repeated for each pair of participants. The accuracy of the classifier was estimated from the proportion of scans correctly classified in both groups and calculated as the average value of sensitivity and specificity. The sensitivity and specificity of the classifier were defined as follows: sensitivity = TP ÷ (TP + FN) and specificity = TN ÷ (TN + FP), where TP = true positives (proportion of images of group 1 correctly classified), TN = true negatives (proportion of images of group 2 correctly classified), FP = false positives (proportion of images of group 2 classified as group 1) and FN = false negatives (proportion of images of group 1 classified as group 2).
Permutation tests
We used permutation testing to derive a p value for the accuracy of each classifier. Here, we permuted the class labels 1000 times (randomly assigning HR or low risk control labels to the training participants) and repeated the cross-validation procedure. We then calculated the number of times in which the specificity and sensitivity for the permuted labels were higher than those obtained for the real labels. Dividing this number by 1000, we derived a p value for the classification accuracies. To estimate the reliability of the 2 methods, we also calculated Cohen κ and the proportion of agreement between SVM and GPC.
Discriminating maps (SVM weight vector)
The use of linear kernel SVM allowed us to directly extract the weight vector as an image (the SVM discrimination map). The SVM decision hyperplane is described by a weight vector and an offset. The weight vector is orthogonal to the hyperplane and corresponds to the most discriminating direction between the groups. Every voxel contributes with a certain weight to the decision boundary or classification function. The SVM weight vector is a linear combination or weighted average of the support vectors and is the spatial representation of the decision boundary. It can be plotted as a brain image to show the relative importance of the voxels in discriminating the classes. Similar to other studies,32 we selected the peaks of the SVM weight vector for each classifier, setting the threshold value to 50% of the maximum (absolute) weight value, and estimated the anatomic regions (cluster peaks) that most contributed to the classifier in the discrimination between groups.
Results
Unaffected HR versus control participants
We compared 45 HR unaffected relatives of BD probands (27 in Halifax, 18 in Prague) to 45 controls without personal or family history of psychiatric disorders who were individually matched by age and sex. All of the unaffected HR participants were medication-naive and medically healthy. The groups were comparable in age, sex, education, handedness and global grey and white matter volumes (Table 1).
Description of the unaffected high risk group and matching controls
Classification accuracy using SVM analysis of white matter images in the combined sample was 68.9% with a sensitivity of 75.6% and specificity of 62.2% (p = 0.001). In other words, among 45 HR participants, 11 individuals were mislabelled as being controls, whereas 17 of 45 controls were incorrectly classified as HR participants. The GPC analysis of white matter images yielded sligtly lower accuracy (65.6%), which was still above chance level (p = 0.002, Table 2). The GPC and SVM showed 96.7% agreement (Cohen κ = 0.93, p < 0.001).
Specificity, sensitivity, accuracy results for discrimination of unaffected participants at high genetic risk for bipolar disorders and low risk controls based on machine learning applied to white and grey matter in each site and in the combined sample
The correctly classified unaffected HR participants were comparable to those misclassified as healthy controls in sex (χ21 = 2.29, p = 0.13), handedness (χ21 = 0.93, p = 0.34) age (t43 = 0.73, p = 0.47), proband diagnosis (BDI v. BDII, χ21 = 0.002, p = 0.96) and proportion of participants with family history of psychosis (χ21 = 0.01, p = 0.91).
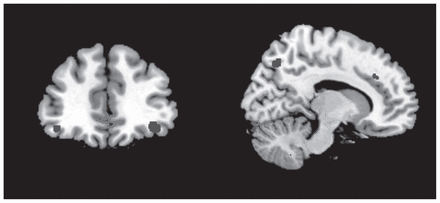
The anatomical regions with the highest contribution to the discrimination of the HR participants from the controls included bilateral white matter tracts adjacent to the ventral prefrontal regions, cingulate gyrus, superior/middle temporal gyrus, precuneus and posterior regions in occipital lobe (Table 3 and Fig. 1).
Regions contributing to discrimination of unaffected participants at high genetic risk for bipolar disorders and low-risk controls included the inferior frontal gyrus (left panel) as well as the cingulate and precuneus (right panel).
Regions contributing to discrimination of unaffected participants at high genetic risk for bipolar disorder and low-risk controls in the combined sample
There were no differences between Halifax and Prague in proportion of BDI probands (74.1% v. 77.8%, χ21 = 0.08, p = 0.78), proportion of participants with family history of psychosis (27.8% v. 26.9%, χ21 = 0.004, p = 0.95), or proportion of female participants (66.7% v. 61.1%, χ21 = 0.29, p = 0.59). The Halifax sample was younger than the Prague group (mean 19.9 v. 21.7 yr, t88 = −2.39, p = 0.019). When we separately analyzed data from the Halifax and Prague samples, the SVM as well as GPC classifiers for white matter also yielded significant classification models with accuracies similar to those in the combined analysis (Table 2). Furthermore, the individual participant SVM weight vectors obtained in the individual site analyses correlated with the weight vectors obtained in the combined sample analyses (r88 = 0.79, p < 0.001). In other words the participants assigned the highest weights in the individual site classifiers also showed the highest weights in the combined sample classifier. The discrimination maps from individual site analyses were mostly congruent with the maps obtained from the combined data analyses (Table 3). When we separately trained the classifiers on data only from Prague or only from Halifax, the resulting discrimination maps mostly overlapped and both contained white matter adjacent to the bilateral inferior frontal gyrus (IFG), middle frontal gyrus (MFG), precuneus, middle temporal gyrus, left fusiform gyrus, left inferior occipital gyrus, right cingulate gyrus, right superior parietal lobule, right inferior temporal lobe and right superior temporal lobe (Table 3).
Machine learning applied to grey matter did not differentiate the 2 groups beyond chance for either the SVM or GPC (Table 2).
Affected offspring versus controls
We compared 36 affected offspring (21 in Halifax, 15 in Prague) to 36 controls without personal or family history of psychiatric disorders, who were individually matched by age and sex. The groups were comparable in age, sex, education, handedness and global grey or white matter volumes (Table 4).
Description of the affected familial participants and matching controls
Classification accuracy using SVM analysis of white matter images in the combined sample was 59.7% with a sensitivity of 58.3% and specificity of 61.1% (p = 0.05). In other words, among 36 affected familial participants, 15 individuals were mislabelled as being controls, whereas 14 of 36 controls were incorrectly classified as affected familial participants. The GPC analyses of white matter images did not discriminate the groups beyond chance.
There were no differences between participants correctly classified as affected and those misclassifed as controls in sex (χ21 = 0.02, p = 0.90), diagnosis (χ25 = 7.71, p = 0.17), handedness (χ21 = 0.06, p = 0.81), treatment (χ21 = 2.62, p = 0.11), age (t34 = 1.09, p = 0.28), duration of illness (t30 = −1.27, p = 0.21), number of episodes (t30 = −0.47, p = 0.64), proband diagnosis (BDI v. BDII, χ21 = 1.07, p = 0.30) or proportion of participants with family history of psychosis (χ21 = 0.01, p = 0.93).
The anatomical regions with the highest contribution to the discrimination of the affected offspring from the controls mostly overlapped with the regions discriminating unaffected from control participants. The maps distinguishing affected from control participants contained more extensive regions, especially in the frontal and parietal lobes, which were not seen in the unaffected versus control comparisons (Table 5).
Regions contributing to discrimination of affected familial participants and low-risk controls in the combined sample
Owing to the smaller sample size of the affected groups, we did not separately analyze the data from Halifax and Prague. Machine learning applied to grey matter did not differentiate the 2 groups beyond chance for either the SVM or GPC.
Discussion
Machine learning applied to structural MRI of white matter was able to discriminate unaffected participants at high genetic risk for BD from healthy low-risk controls above the chance level, with an accuracy of 68.9%. In addition, 2 different methods of ML analyses, GPC and SVM, showed 96.7% agreement. The accuracy decreased to 59.7% when we attempted to differentiate the more clinically heterogeneous affected familial participants from healthy controls.
No previous brain imaging studies have used ML in unaffected offspring of parents with BD. The 68.9% accuracy of classification has good face validity. A proportion of unaffected offspring of parents with BD may not have inherited biological risk factors for BD and may not differ from controls in their brain structure. Thus, higher classification accuracy would suggest overfitting.
The results for the affected participants are comparable to previous literature. Similar to our findings, Schnack and colleagues38 also reported 59% accuracy when distinguishing participants with BD from healthy controls. The lower classification accuracy among the affected than the unaffected groups may reflect confounding by clinical variables. There is replicated evidence suggesting that repeated episodes of illness,8,9 comorbid conditions,13,14 or exposure to medications10–12 introduce heterogeneity, which may mask/overcome the primary changes indicative of the risk for BD.16,18,19 The findings emphasize the need to control for clinical heterogeneity, for example by recruiting unaffected participants at high genetic risk for the illness.
Interestingly, a single previous ML study using GPC of white matter was able to differentiate participants with established BD from controls with accuracies between 69% and 78%.30 These higher accuracies may reflect differences between the studies in relevant clinical variables. Some of the brain alterations in participants with BD may accumulate with illness burden. It is thus relevant that, relative to our sample or even that of Schnack and colleagues,38 Rocha-Rego and colleagues30 recruited participants with longer duration of illness, in whom the secondary, neuroprogressive changes may have been more pronounced and could have contributed to the differentiation from healthy controls. Indeed, a previous study has shown that among participants with psychotic episodes, ML was sensitive to differences in illness course.32
Because the SVM classifiers are based on the whole brain patterns and take into account spatial correlations in the data, it is difficult to make local inferences based on these approaches. Nevertheless, the regions that most contributed to the distinction between unaffected HR, affected familial and healthy control participants have previously been detected as abnormal in patients at risk for the illness18 or those with fully manifest BD.41 Notably, the discrimination maps included the right inferior frontal gyrus, which in our previous voxel-based morphometry study in an overlapping sample showed replicated differences between both unaffected and affected familial participants and controls.18 The fact that 2 very different methods of data analysis identified structural alterations in the same region as a potential biomarker of BD is encouraging and supports the biological validity of this finding. Notably, when we separately trained the classifiers on data only from Prague or only from Halifax, the resulting discrimination maps mostly overlapped. The algorithm was able to distinguish the HR from control participants based on similar global neuroanatomical patterns of white matter changes.
The maps differentiating unaffected or affected participants from controls contained some of the same regions (Table 3 and Table 5). These overlapping regions/networks may be associated with susceptibility for BD rather than with resilience. It is also of note that the affected participants showed additional changes that may be secondary to illness-related variables. In keeping with this, the volume of regions uniquely contributing to differentiation between affected and control groups (i.e., the superior and medial frontal gyrus, the supramarginal gyrus and the other parietal lobe regions) has in previous studies been associated with unique, BD-related variables more than with shared genetic factors.19
The neuroanatomical maps that discriminated the HR or affected familial participants from controls were distributed throughout the whole brain rather than highly localized. This is in keeping with other studies and neuroanatomical models of psychiatric disorders.1,5 The maps included regions involved in voluntary or automatic emotion regulation (right inferior frontal gyrus, ventrolateral prefrontal cortex, cingulate), attention/executive functions (prefrontal regions) and self-monitoring (precuneus and other parietal cortex regions),42–44 which are relevant to the pathophysiology of BD.42,43,45 Many of the same regions have also been reported to be abnormal in individuals with unipolar depression or schizophrenia.46,47 We would need groups of patients with other disorders to assess the specificity of these discriminating patterns. Interestingly, a previous ML study using structural MRI was able to accurately and significantly differentiate patients with BD from participants with schizophrenia, but not from healthy controls.38
We were not able to distinguish healthy participants at high genetic risk for BD or affected offspring from healthy low-risk controls based on grey matter structure. This is congruent with another study in which SVM of grey matter distinguished BD from control participants with an accuracy of 59% and sensitivity of 53%38 as compared with 57% accuracy and 53% sensitivity in our study.
Limitations:
Our study has the following limitations. Machine learning studies benefit from large sample sizes. The predictive models become stable at about 130 participants per group.48 We included a total of 126 participants. For comparison, a previous study of BD investigated 80 participants,30 and the first ML study in participants at clinical risk for schizophrenia included 70 individuals.31 In addition, our study benefited from the availability of 2 independent samples from 2 sites, which functioned as replication cohorts. Also, the results fit closely with those of previous studies of patients with BD or those at risk for the disorder and show high biological validity. One of the concerns with ML is the potential for overfitting of models. We used conservative methods of cross-validations, similar to previous studies.30,32,39 In addition, we separately analyzed participants from 2 cohorts, which yielded similar accuracies and overlapping discrimination maps. Finally, it is unlikely that our results represent overfitting, as none of the grey matter–based classifiers was able to differentiate the 2 groups.
Although the groups were comparable in handedness, numerically there were more left-handed participants in the unaffected HR group. We did not have measures of IQ or across-sites interrater reliability for the psychiatric diagnoses. However, the groups were comparable in education levels. In each site we used state-of-the-art diagnostic procedures, including assessments by board-certified psychiatrists, access to clinical information from prospective follow up and blind consensus meetings by panels of senior clinical researchers. In addition, 2 of us (T.H., M.A.) have appointments at both institutions, and 1 (T.H.) has worked clinically in both centres, which further facilitates consistency of the assessments.
The main advantage of this study was the use of the HR design. Clinical heterogeneity with regards to medication exposure, duration of illness and comorbid conditions influences brain structure in individuals with BD, sometimes in opposing directions.8,18 The fact that the unaffected groups included only medication-naive, unaffected participants without any comorbid conditions makes the interpretation of findings much easier. Although the HR design decreases the clinical heterogeneity, it may introduce other sources of heterogeneity vis-à-vis resilience/protective factors. The addition of the affected familial group allowed for a better interpretation of findings. We recruited participants in the age range when transition to BD is most likely23,24 and when the diagnostic use of neuroimaging would be most useful. The accuracy of the ML classifications needs to be determined against “gold standard” diagnostic assessments. It is therefore important that the probands received a detailed psychiatric interview conducted by psychiatrists and that their diagnosis was established based on consensus of the research group. Finally, we used a cutting-edge technique for MRI data analyses, which could help realize the diagnostic potential of MRI in psychiatry.
Conclusion:
In this study, ML combined with structural MRI was able to discriminate healthy participants at high genetic risk for BD from healthy low-risk controls above the chance level, with an accuracy of 68%. Our findings suggest that distributed patterns of white matter changes may be of greater diagnostic utility for early detection of BD than grey matter biomarkers. The discrimination maps included some of the main candidates for biological risk factors of BD and had a good face validity vis-à-vis the functional neuroanatomy of BD. These results provide a proof of concept that neuroimaging could potentially contribute to early identification of individuals with BD or those at risk for the disorder. Once we have a library of sufficiently robust ML kernels for individual disorders or specific stages of illness, we may be able to compare the MRI of new study participants against this library to help with the diagnostic process. Addition of genetic and biochemical data could further improve classification accuracy.
Acknowledgements
This study was supported by funding from the Canadian Institutes of Health Research (103703, 106469), the Nova Scotia Health Research Foundation and grant from the Ministry of Health (NT13891) of Czech Republic. The sponsors of the study had no role in the design or conduct of this study; in the collection, management, analysis, and interpretation of the data; or in the preparation, review, or approval of the manuscript. The authors report no biomedical financial interests or potential conflicts of interest.
Footnotes
Competing interests: The authors are supported by grants from the Canadian Institutes of Health Research (103703, 106469), the Nova Scotia Health Research Foundation and the Ministry of Health of Czech Republic (NT13891). C. Höschl declares personal fees from Servier, Lundbeck International Neuroscience Foundation, Eli Lily and Janssen-Cilag. No other competing interests declared.
Contributors: T. Hajek and M. Alda designed the study. T. Hajek, M. Kopecek, T. Novak and M. Alda acquired the data, which T. Hajek, C. Cooke and C. Höschl analyzed. T. Hajek wrote the article, which all authors reviewed and approved for publication.
- Received May 29, 2014.
- Revision received October 16, 2014.
- Accepted December 23, 2014.