Abstract
Background: Among healthy participants, the interindividual variability of brain response to facial emotions is associated with genetic variation, including common risk variants for schizophrenia, a heritable brain disorder characterized by anomalies in emotion processing. We aimed to identify genetic variants associated with heritable brain activity during processing of facial emotions among healthy participants and to explore the impact of these identified variants among patients with schizophrenia.
Methods: We conducted a data-driven stepwise study including samples of healthy twins, unrelated healthy participants and patients with schizophrenia. Participants approached or avoided pictures of faces with negative emotional valence during functional magnetic resonance imaging (fMRI).
Results: We investigated 3 samples of healthy participants — including 28 healthy twin pairs, 289 unrelated healthy participants (genome-wide association study [GWAS] discovery sample) and 90 unrelated healthy participants (replication sample) — and 1 sample of 48 patients with schizophrenia. Among healthy twins, we identified the amygdala as the brain region with the highest heritability during processing of angry faces (heritability estimate 0.54, p < 0.001). Subsequent GWAS in both discovery and replication samples of healthy non-twins indicated that amygdala activity was associated with a polymorphism in the miR-137 locus (rs1198575), a micro-RNA strongly involved in risk for schizophrenia. A significant effect in the same direction was found among patients with schizophrenia (p = 0.03).
Limitations: The limited sample size available for GWAS analyses may require further replication of results.
Conclusion: Our data-driven approach shows preliminary evidence that amygdala activity, as evaluated with our task, is heritable. Our genetic associations preliminarily suggest a role for miR-137 in brain activity during explicit processing of facial emotions among healthy participants and patients with schizophrenia, pointing to the amygdala as a brain region whose activity is related to miR-137.
Introduction
Facial expressions of primary emotions are universally recognized by humans and are considered innate.1,2 They carry crucial information about what to expect from social interactions and, more generally, from the environment. Recognizing facial expressions that signal threats, such as angry and fearful faces, can be important to avoid harm and deal with potentially dangerous situations.3
Visual processing of angry and fearful faces is associated with activity in several brain regions, including the limbic areas (i.e., amygdala and parahippocampal gyrus, posterior cingulate cortex), temporoparietal areas (i.e., inferior and superior parietal lobule, middle temporal gyrus), insula, prefrontal areas (i.e., inferior and medial frontal gyrus), subcortical areas (i.e., putamen) and the cerebellum,4 as revealed by functional neuroimaging. In particular, activity in the prefrontal areas and amygdala is greater during explicit than implicit paradigms, when the emotional content of faces is explicitly evaluated and not just automatically processed at a more basic level of consciousness.5 This finding reflects a direct involvement of the prefrontal cortex and amygdala in processing threatening and social emotion stimuli.
However, although compelling evidence from behavioural research on infants, blind people and twins have consistently supported an innate source for expression and recognition of facial emotions,6,7 the cerebral and biological evidence toward the heritability of this behaviour remains sparse. Recent imaging genetics studies have shown that activity in brain areas specifically involved in response to threatening faces is associated with common genetic variants.8 For example, the combination of functional imaging paradigms and single-gene data has yielded associations between prefrontal and limbic brain activity with genetic variants of the SLC6A4 serotonin transporter,9 the catechol-O-methyltransferase enzyme (COMT gene),10 the monoamine oxidase A enzyme (MAOA gene),11 and the dopamine D2 receptor (DRD2 gene).12,13
Notably, previous studies were based on hypotheses about specific brain regions or specific genetic variants. Thus far, only few variants have been associated with processing of facial emotions. In a data-driven neuroimaging genetics study, Elliot and colleagues,14 using data from the UK Biobank, searched for genome-wide association with 3144 functional and structural brain phenotypes, finding no significant association with task-based phenotypes on functional magnetic resonance imaging (fMRI), including implicit processing of facial emotion. However, this study considered brain activity of imaging-derived phenotypes as a whole, summarizing measures of functional activity in response to different tasks (thus, also nonemotional tasks). It is possible that identifying common genetic variants related to the brain’s underpinnings of facial emotion processing requires more specific hypotheses about the brain regions involved and the details of the tasks used to probe brain activity. Moreover, knowledge about heritability of brain regions’ activity during processing of facial emotions, which can be investigated in twins, is wanting. Indeed, we contend that a brain map of such heritability would validate the focus on brain areas whose function is most strongly associated with genetic variation, thus increasing the statistical power of the investigated association and preserving the functional specificity of the brain regions.
The relevance of this line of research is evident when considering that severe psychiatric disorders often include symptoms of emotion dysregulation.15 For example, the diagnosis of schizophrenia includes affective flattening and dysregulated social behaviour on the one hand and impaired perception of facial expressions on the other.16 These impairments are core domains of social cognitive dysfunction in schizophrenia, are linked to psychopathology and are predictors of functional outcomes.17 A growing number of studies involving patients with schizophrenia shows that impaired facial perception is detected, especially for threatening facial expressions such as fearful and angry faces.18–21 Moreover, dysregulation of approach and avoidance (i.e., fight or flight) behaviour is a key aspect of social dysfunction in schizophrenia (e.g., increased emotionality but deficit in perception and expression of emotion).17,22,23
The diagnosis of schizophrenia is highly heritable.24 Notably, the third multistage genome-wide association study (GWAS), conducted by the Psychiatric Genomic Consortium (PGC), identified 287 independent genetic loci conferring risk for schizophrenia.25 Supporting the idea that genetic risk for schizophrenia is also associated with emotion processing in unaffected people, Germine and colleagues26 showed that the polygenic risk score for schizophrenia, cumulating the effects of many polymorphisms, is associated with higher reaction times when recognizing facial emotions among children, starting already at age 9 years. However, a recent study involving healthy adults found no significant association between the polygenic risk score for schizophrenia and activity in any brain region.27 The analysis of separate single nucleotide polymorphisms (SNPs) did reveal that rs9607782 — located near the gene EP300, which codes for p300, a protein involved in fear memory consolidation and amygdala plasticity — was significantly associated with amygdala recruitment during implicit processing of angry and fearful faces. Furthermore, rs1702294, a SNP located in miR-137 — a post-transcriptional master regulator and one of the most strongly associated loci for schizophrenia risk (p = 6.6 × 10−16)28 — was nominally associated with right amygdala activity but did not survive correction for the number of emotion-related regions of interest, selected a priori. However, the sample size and characteristics (245 participants out of 578 presented with family history of major psychiatric disorders), as well as the hypothesis-based selection of SNPs and regions of interest, may have limited this investigation. Together, these studies suggest that composite genetic risk for schizophrenia includes variants that play a role in emotion processing, but pinning down these effects to specific brain regions is more difficult. In addition, this study supports a nominally significant role of the miR-137 variant in amygdala activation during implicit processing of facial emotions, and not the genetic underpinnings of explicit processing, which includes decision-making in response to emotion recognition.
Thus, the main aim of the present study was to identify, through a data-driven stepwise approach, common genetic variants associated with brain activity during the processing of faces with negative valence. In the first step, we aimed to evaluate the heritability of whole brain activity among twins during the processing of facial stimuli of fear and anger, in a paradigm eliciting explicit social evaluation of facial expressions. In the second step, we aimed to evaluate the association of common genetic variants with the brain areas whose activity was heritable, using a discovery sample and a replication sample of healthy unrelated participants. Finally, given the findings on genetic variation associated with the heritable brain activity to facial emotions that was also involved in risk for schizophrenia, we further sought to explore the impact of genetic variation in a third independent data set of patients with schizophrenia.
Considering that facial expressions of fear and anger are universally recognized and innate, that the prefrontal cortex and the amygdala are markedly involved in attention to threat and social evaluation of facial expressions and that imaging genetic studies using single-gene approaches support the hypothesis that genetic variation contributes to individual differences in the activity within the brain circuit involved in processing facial emotions, especially threatening faces, we hypothesized that activity of the prefrontal and amygdala regions in response to explicit processing of fearful and angry faces is heritable. Importantly, we expected that investigating the genome-wide association of common variants with heritable activity of specific brain regions would enhance the detection of replicable associations. Our overarching aim was a proof of concept on the genetic architecture of emotion dysregulation, which is a key symptom of many highly heritable brain disorders, including schizophrenia.
Methods
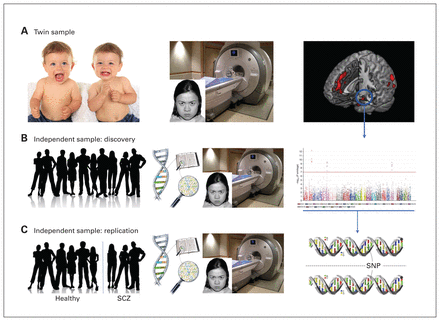
Using a data-driven stepwise approach, we designed an exploratory study to identify common genetic variants associated with brain activity during the processing of faces with negative emotional valence (Figure 1).
Study design. We used a data-driven multi-step approach in which we first detected the brain region whose activity during explicit processing of threatening faces had the highest heritability, investigating (A) a twin sample of healthy individuals. (B) Subsequently, we conducted a genome-wide association study meta-analysis on that specific regional activity using independent discovery and replication samples of healthy non-twins. (C) Finally, we sought replication in a sample of patients with schizophrenia (SCZ).
Participants
We recruited participants from Apulia, Italy, divided into 3 samples of healthy participants and 1 sample of patients with schizophrenia. The first 3 groups included a sample of healthy twin pairs, a sample of unrelated healthy participants for a GWAS discovery sample and a sample of unrelated healthy participants as the first replication sample, in which a different batch composition was used for genotyping. We used a sample of patients with schizophrenia as the second replication sample. We used the Structured Clinical Interview for DSM-IV to confirm the diagnosis of schizophrenia for patients and to exclude any Axis I psychiatric disorder for healthy participants.
Zygosity estimations
Among twins, we inferred zygosity using a standardized questionnaire (2 peas in a pod; 95% accuracy29), administered to parents. We used twin samples, including monozygotic and dizygotic twin pairs, for heritability estimation of emotion-related brain activity.
Neuropsychological task
During fMRI, we presented angry, fearful, happy and neutral facial expressions from a validated set of facial pictures (NimStim, http://www.macbrain.org/resources.htm). We instructed participants to decide if they would like to approach or avoid the face (explicit facial emotion processing). For this study, we focused on the analysis of angry and fearful faces (Appendix 1, available at www.jpn.ca/lookup/doi/10.1503/jpn.230013/tab-related-content, for additional information). Most healthy participants in this study also conducted a second task, involving the implicit processing of facial emotion of the same faces.
Functional magnetic resonance imaging data acquisition and analysis
We performed blood oxygen level–dependent (BOLD) fMRI on a GE Signa 3T scanner while participants performed the emotion task. We analyzed fMRI data using Statistical Parametric Mapping 8 (Wellcome Department of Cognitive Neurology). Appendix 1 includes details on data acquisition and preprocessing. We entered individual contrast images in the intraclass correlation (ICC) analyses,30 as well as in a second-level random effects analysis. We performed separate t tests of angry versus baseline and fearful versus baseline (regardless of the approach or avoidance choice) on all the samples used in this study. We then extracted activity in these contrasts within the regions of interest selected via the ICC analyses using the MarsBar toolbox (http://marsbar.sourceforge.net/). Given that our aim was to characterize brain area activation during processing of threatening faces and that processing of neutral faces activates mostly the same brain areas as of threatening faces,4,31,32 the neutral condition represents a contentious baseline, as mentioned elsewhere.33,34 Therefore, in line with our study aims and coherently with several previous studies,4 we used a fixation crosshair as a baseline to control for activity related to basic body functions (e.g., breath, basic vision, space perception, body temperature regulation) while preserving detection of brain activity during processing of facial emotions (more details are available in Appendix 1).
Estimates of genetic contribution on brain activity during processing of threatening face emotions
To identify areas with heritable activity during processing of threatening facial emotions, we selected the brain areas in which the activity was highly correlated with the whole brain activity of angry and fearful faces, as per the ICC analysis of data from monozygotic twins (p, corrected for family-wise error [FWE] < 0.05; ICC > 0.6) (Appendix 1).30 We jointly estimated genetic and environmental contributions of these areas by fitting an ACE model (where A is the additive genetic, C is the common familial environment and E is the environmental contribution unique to the individual), which considers the covariance of monozygotic and dizygotic twins, as well as the total phenotypic variance (Appendix 1).
We extracted the BOLD signal from the brain areas with significant heritable activity and conducted a GWAS of healthy participants to identify genetic variants associated with the BOLD parameter estimates of these areas.
Genome-wide association study
Genotyping
Healthy participants and patients with schizophrenia underwent blood withdrawal for subsequent DNA extraction from peripheral blood mononuclear cells. We genotyped 2 groups of healthy participants using the same procedure.35 We used the largest group as a discovery set and the other as an independent replication set. Details of GWAS procedures are shown in Appendix 1.
Quantitative trait loci analysis
We used the BOLD signal extracted from amygdala regions of interest (generated using voxels with heritable activity) from both the discovery and healthy replication samples. The GWAS threshold for a discovery sample is usually set to a p value less than 5 × 10−8, with a 1-tailed p value less than 0.05 for the replication sample. However, our sample size was limited for a GWAS,36,37 and the discovery sample was underpowered to detect common variants below the usual threshold. Thus, we further meta-analyzed p values of the same SNP from discovery and replication samples and computed an alternative, less stringent genome-wide threshold.
Meta-analysis of genome-wide association study
We used the Stouffer formula to combine the p values of the same SNP, weighted by the square root of the sample sizes of the discovery and replication samples.38 The Stouffer method is often used in meta-analysis to combine p values of genetic variants across independent studies, including discovery and replication studies, and derive an overall significance.39–41 This approach allows for the integration of findings from multiple cohorts or populations, increasing statistical power and providing a more comprehensive understanding of the genetic associations with complex traits or diseases.42–44 The Stouffer method is implemented in the metap R package (sumz function). We retained only SNPs with a concordant allelic dosage effect (i.e., SNPs whose reference allele is associated with reduced or increased BOLD signal in both discovery and replication sets).
Alternative thresholding for multiple tests
Given that the usual GWAS threshold may be too stringent for our limited sample size, diluting true positive detection,36,37 we used an alternative method to estimate the number of independent tests within our sample,45 which is based on the principal component analysis decomposition of genotype matrices. Indeed, the standard thresholds of significance in GWAS are generally considered to be p less than 5 × 10−8 or p less than 1 × 10−8, for type I α levels of 0.05 and 0.01, respectively.46 However, the restraint of type I error may inflate type II error because the standard GWAS method is based on a Bonferroni correction for an assumed million independent genetic variants in the human genome. In studies with low power or low-frequency alleles, a Bonferroni correction has been shown to be overly conservative by detrimentally inflating the type II error.47 Alternative correction methods can ameliorate the loss of power owing to a small sample size, when gathering large numbers of participants is impractical.45,47 Thus, we used the SimpleM software as an alternative method of correcting for multiple testing. SimpleM encompasses 3 steps, namely computation of the composite linkage dis-equilibrium correlation matrix from the SNP data set, calculation of the eigenvalues and calculation of the effective number of independent tests through principal component analysis, which amounted to 227 319 independent tests. Thus, the threshold p value of the 5% genome-wide significance was adjusted to 2.2 × 10−7 (0.05/227 319).
Results
Participants
Our final study sample included 483 White participants, divided into 3 samples of healthy participants and 1 sample of patients with schizophrenia, including 28 healthy twin pairs (16 monozygotic and 12 dizygotic twin pairs), 289 healthy participants for the GWAS discovery sample, 90 healthy participants for the first replication sample and 48 patients with schizophrenia for the second replication sample (Table 1, with further details on participant demographics reported in Appendix 1). In the first replication sample, 1 individual was excluded as an outlier on brain activity estimates (2-tailed Grubbs test p < 0.05), with 89 participants remaining. Of the total sample, 315 healthy participants also conducted a second neuroimaging task, involving the implicit facial emotion processing. Analyses and results of this additional task run are shown in Appendix 1.
Participant demographics
Neuroimaging results
Results of t tests from the 4 group samples of this study revealed significant activation (FWE-corrected pFWE < 0.05) in brain areas during processing of facial emotions for the angry versus baseline and fearful versus baseline conditions (i.e., bilateral superior, middle and inferior frontal gyrus; bilateral thalamus and hippocampus; para-hippocampus; amygdala; bilateral fusiform gyrus, lingual gyrus and posterior and anterior cingulate gyrus). Moreover, these maps served for the extraction of BOLD estimates from the clusters selected via ICC analyses. We entered these extracted values in the ACE model and GWAS analyses.
Heritability results
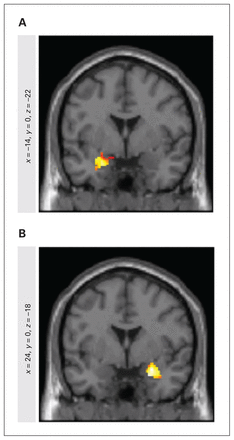
Among healthy twins, we identified the bilateral amygdala as the brain region with the greatest heritability during explicit processing of angry faces, as evaluated with our fMRI task (right amygdala: x = 24, y = 0, z = −18; ICC = 0.83, κ = 18, p < 0.001; left amygdala: x = −14, y = 0, z = −22; ICC = 0.66, κ = 5, p < 0.001) (Figure 2). Since we did not find significant differences between the ICC values of the left and right amygdala (t test p > 0.05), we combined the signals of these 2 regions, and recalculated the ICC (ICC = 0.79, FWE-corrrected p < 0.05). After fitting the ACE model, we obtained a heritability estimate (a2) of 0.54 (common environment c2 = 0.4, unique environment e2 = 0.06).
Functional magnetic resonance images, showing (A) left and (B) right amygdala activity in the twin sample during processing of face emotions. The combined signals of these 2 regions revealed an intraclass correlation of 0.79 (p < 0.001) and a heritability estimation (a2) of 0.54.
We did not find any statistically significant ICCs in brain activity among monozygotic twins during the explicit processing of fearful faces.
To avoid type II errors, we also explored ICC data at alternative thresholds of an ICC greater than 0.5 and an uncorrected p value less than 0.005, and conducted subsequent GWAS analyses (the results of these explorations are shown in Appendix 1). Here, we report the results obtained with the most stringent threshold used (ICC > 0.6).
Genome-wide association study results
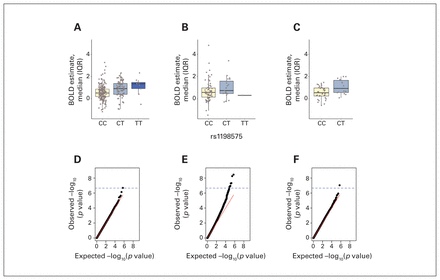
The subsequent GWAS, involving healthy non-twins, indicated that bilateral amygdala activity during the emotion processing task was associated with the rs1198575 polymorphism (p = 1.5 × 10−7) (Figure 3). This result survived the SimpleM corrected threshold for multiple testing (2.2 × 10−7) (Appendix 1, Figure 1). This SNP is located in the 5′ region of the miR-137 host gene, at 51 kilobase pairs from miR-137 and at 47 kilobase pairs from the host gene. Specifically, the C allele is associated with lower amygdala activity during recognition of angry faces, compared with the T allele (Figure 3A). The genotypic effect of this SNP was significant in the same direction in the replication sample (p = 0.01) (Figure 3B) and among patients with schizophrenia (p = 0.03) (Figure 3C).
Quantitative trait locus analyses of rs1198575 and quantile–quantile plots. Panels A through C show boxplots depicting the robust linear additive effect of the T allele on blood oxygen level–dependent (BOLD) estimates in the amygdala among (A) healthy participants (discovery sample), (B) healthy participants (replication sample) and (C) patients with schizophrenia (replication sample). Horizontal thick lines show medians. Boxes show interquartile ranges (IQRs). Vertical lines show the largest values within 1.5 times the IQR above or below the third or first quartile, respectively. The lower panels show p value distributions for genome-wide association analyses of BOLD estimates in the amygdala for (D) healthy participants (discovery sample), (E) healthy participants (replication sample) and (F) healthy participants with the combined p value using the Stouffer formula. The horizontal dashed blue lines show the SimpleM threshold for genome-wide statistical significance (2.2 × 10−7).
The GWAS meta-analysis in discovery (p = 1.5 × 10−7) and replication (p = 0.01) samples of healthy non-twins indicated that bilateral amygdala activity during the task was associated with the polymorphism rs1198575 (p = 9.1 × 10−8), as shown in Figure 3F. Table 2 and Appendix 1, Table 1 show detailed GWAS results.
Results of the genome-wide association meta-analysis*
Discussion
This study explored the genome-wide association of common genetic variations with brain activity during explicit processing of faces with negative valence (i.e., angry and fearful faces). To this aim, we used a data-driven multistep approach in which we first detected the brain regions whose activity during processing of threatening faces had the highest heritability; subsequently, we conducted a GWAS meta-analysis on brain regional activity using independent discovery and replication samples of healthy non-twins; finally, we sought replication in a sample of patients with schizophrenia. The results from the small sample of twins suggested that amygdala reactivity to judgment of angry faces (i.e., approach or avoid responses) is significantly heritable. Furthermore, our results from a healthy non-twin sample suggested that a polymorphism close to miR-137 (rs1198575) has a genome-wide association with amygdala response during processing of angry faces, with C-allele carriers having lower activity than participants who carried TT alleles. This finding was also replicated in an independent sample of healthy participants and in a sample of patients with schizophrenia. Previous findings have shown that this polymorphism has a genome-wide association with schizophrenia,28 with the C allele conferring risk for the disorder.
The amygdala is a key region for the processing of facial emotions5 and its volume, as measured with structural MRI, is significantly heritable.48 Our twin study, which used fMRI to investigate heritability of brain activity during explicit processing of face emotions, suggests that amygdala activity related to face emotions is significantly heritable, as previously suggested by single-gene candidate studies.12,13 Alterations of amygdala activity during processing of facial emotions have been consistently found in several psychiatric disorders, including schizophrenia.49 Specifically, a recent meta-analysis found significant reductions of amygdala activity among patients with schizophrenia during processing of faces with negative valence.50 Also, consistent with our study, previous findings have indicated amygdala hypoactivity among people at genetic risk for psychiatric disorders.51
MicroRNAs are small noncoding RNA molecules that regulate the expression of some target genes at a post-transcriptional level,52 and can modulate several neuronal and behavioural phenotypes.53 Specifically, many studies indicate that miR-137, a brain-enriched microRNA, is critical for the life and activity of neural cells in both developing and mature brains.54 Specifically, it regulates the balance between proliferation and differentiation of neurons by inhibiting the first process and promoting the second process.54 Recently, He and colleagues55 identified miR-137 as an important regulator of synaptic function in the adult brain. This finding is particularly remarkable for our study, since alterations in synaptic features (including decreased synaptic connectivity and transcriptional dysregulation of certain genes) are frequently found in psychiatric disorders56 and genetic risk for schizophrenia is thought to converge on synaptic biology.25 Indeed, Siegert and colleagues57 have investigated the functional impact of psychiatric risk of SNPs in miR-137. They observed increased miR-137 levels in induced human neurons harbouring the minor alleles of 4 schizophrenia-associated SNPs in miR-137, compared with major (most common) allele–carrying cells. More specifically, Siegert and colleagues57 propose a model through which a single regulatory microRNA could simultaneously control several protein targets, thus regulating an entire pathway, resulting in synaptic alterations. Moreover, our results are consistent with findings showing the amygdala as one of the brain regions with the highest levels of miR-137 expression.58
In addition, a variant in miR-137 has been previously linked with frontoamygdala emotion connectivity among healthy participants.59 Our results are consistent with these findings and further highlight a crucial role for miR-137 in emotion dysregulation. On the other hand, our findings diverge from those found in the GWAS of Elliot and colleagues.14 However, this study investigated the genetic associations related to implicit — not specifically explicit — face processing, which requires a stronger engagement of the amygdala and prefrontal cortex.4 Notably, Elliot and colleagues14 combined fearful and angry faces, which may be related to different brain responses,60 and investigated the heritability of participant-specific activation of imaging-derived phenotypes instead of voxel-wise data, thus aggregating brain functional response to different tasks (including cognitive tasks).
The replication of our GWAS association in a sample of patients with schizophrenia corroborates the role of miR-137 in emotion dysregulation. Interestingly, in the first PGC GWAS study, Ripke and colleagues61 identified miR-137 (rs1625579) as the locus with the strongest association with schizophrenia (p = 1.6 × 10−11). They also proposed a subset of the 4 genes targeted by miR-137 that were associated with schizophrenia.61 These results were replicated in the second PGC study, which showed that the locus with the second strongest correlation with schizophrenia was within an intron of miR-137 (p = 3.4 × 10−19).28 Moreover, Yao and colleagues62 used data from the third PGC study and found that polygenic risk score derived from sets of target genes of miR-137 explained a disproportionately larger variance in schizophrenia risk than genome-wide SNP sets. However, these studies could not shed light on the functional role of this gene in the context of emotion-related phenotypes. On the other hand, our study, although using a small sample for GWAS analysis, features a paradigm designed to index brain activity specific to explicit processing of face emotions. This procedure allowed us to search for subtle functional effects of genetic variants on the brain. This approach showed that a polymorphism close to miR-137 (and close to the polymorphism found by Erk and colleagues27) is the only one that survived correction for multiple testing and was associated with functional brain activity during processing of face emotions among both healthy participants and in patients with schizophrenia. This preliminary evidence suggests that, among all the genetic variants associated with risk of schizophrenia, miR-137 harbours variants that are biologically plausible candidates related to dys-regulations in processing of face emotions in this disorder.
We did not find any statistically significant ICCs in brain activity among monozygotic twins during the explicit processing of fearful faces. A speculative interpretation of these results may rely on evidence that angry and fearful faces, although both negative stimuli, represent qualitatively different forms of threat. Fearful faces are thought to signal an undetermined, ambiguous threat, while angry faces represent a more direct form of threat, often used in face-to-face encounters to exert dominance.60,63 According to this view, the brain response associated with the approach or avoidance of an ambiguous threat, such as fearful faces, may be modulated by learned experience, while the brain response associated with the approach or avoidance of a direct threat, such as angry faces, may be more related to instinctive, heritable response patterns involving the amygdala.
Limitations
Our sample size was small for a GWAS analysis. However, we tried to mitigate this limitation in different ways via a data-driven approach (i.e., heritability estimation of whole brain activity). We selected the brain signals for which we would test GWAS associations, thus focusing on a heritable phenotype. Moreover, we conducted a meta-analysis on p values of the same SNPs from discovery and replication samples and computed an alternative correction for multiple testing based on principal component decomposition, which is less dependent on the sample size, instead of composite linkage dis-equilibrium values.45 We used a second replication data set of patients with schizophrenia to further validate the genetic association. All of these procedures are meant to limit the probability of type I and type II errors. We did not find a similar data set for external replication. This is owing to the uniqueness of our fMRI task. Indeed, although the emotion fMRI paradigms are used worldwide, the explicit emotion paradigm involving social evaluation of negative faces is less common than the implicit paradigm. Thus, despite the small sample size, this study is a proof of concept that miR-137 risk variants for schizophrenia may underpin explicit face emotion processing among neurotypical participants. Further studies with larger schizophrenia samples are necessary to directly explore the potential association between the rs1198575 polymorphism and diagnosis on the amygdala activation during facial emotion processing. We calculated zygosity via a questionnaire and not genetic analysis. The questionnaire used has an accuracy of 95% (compared with 99% accuracy of genetic analysis), exposing our analyses to possible misclassification of 1 out of 28 pairs, on average. We believe that this is a relatively small risk.
Conclusion
Our study provides a heritability map of brain activity related to processing of angry faces, identifying the bilateral amygdala as the region with the greatest heritable activity. It also provides a GWAS association from functional imaging data, highlighting a role for miR-137 in explicit processing of facial emotions and its dysregulation.
Footnotes
↵* These authors contributed equally to this work.
Competing interests: Giuseppe Blasi received lecture fees by Lundbeck. Marco Papalino received travel fees by Newron Pharmaceuticals. Alessandro Bertolino reports speaker fees from Lundbeck, Otsuka and Angelini. No other competing interests were declared.
Contributors: Tiziana Quarto, Giuseppe Blasi, Giulio Pergola and Alessandro Bertolino contributed to the conception and design of the work. Annalisa Lella, Pasquale Di Carlo, Antonio Rampino, Vittoria Paladini, Marco Papalino, Raffaella Romano, Leonardo Fazio, Daniela Marvulli and Teresa Popolizio contributed to data acquisition, analysis and interpretation. Tiziana Quarto, Annalisa Lella, Giuseppe Blasi, Giulio Pergola and Alessandro Bertolino drafted the manuscript. All of the authors revised it critically for important intellectual content, gave final approval of the version to be published and agreed to be accountable for all aspects of the work.
Funding: This project received funding from the European Union Seventh Framework Programme for research, technological development and demonstration (grant no. 602450). Annalisa Lella was supported by a grant (no. 2017M7SZM8 [PRIN2017]) awarded by Alessandro Bertolino and by research funding from Apulia regional government to Alessandro Bertolino (DGR no. 1702/2020). Giulio Pergola was supported by the European Union’s Horizon 2020 research and innovation program under the Marie Sklodowska-Curie grant (no. 798181 [FLOURISH]). Antonio Rampino, Giulio Pergola and Alessandro Bertolino were funded by the European Union funding within the Ministero dell’Università e della Ricerca PNRR extended partnership initiative on neuroscience and neuropharmacology (no. PE00000006 CUP H93C22000660006). Alessandro Bertolino was funded by the Ministry of Economic Development (Project no. F/200044/01-03/X45). This article reflects only the authors’ views and the European Union is not liable for any use that may be made of the information contained therein.
- Received January 16, 2023.
- Revision received July 11, 2023.
- Accepted July 30, 2023.
This is an Open Access article distributed in accordance with the terms of the Creative Commons Attribution (CC BY-NC-ND 4.0) licence, which permits use, distribution and reproduction in any medium, provided that the original publication is properly cited, the use is noncommercial (i.e., research or educational use), and no modifications or adaptations are made. See: https://creativecommons.org/licenses/by-nc-nd/4.0/