Abstract
Studies of clinical populations that combine MRI data generated at multiple sites are increasingly common. The Canadian Biomarker Integration Network in Depression (CAN-BIND; www.canbind.ca) is a national depression research program that includes multimodal neuroimaging collected at several sites across Canada. The purpose of the current paper is to provide detailed information on the imaging protocols used in a number of CAN-BIND studies. The CAN-BIND program implemented a series of platform-specific MRI protocols, including a suite of prescribed structural and functional MRI sequences supported by real-time monitoring for adherence and quality control. The imaging data are retained in an established informatics and databasing platform. Approximately 1300 participants are being recruited, including almost 1000 with depression. These include participants treated with antidepressant medications, transcranial magnetic stimulation, cognitive behavioural therapy and cognitive remediation therapy. Our ability to analyze the large number of imaging variables available may be limited by the sample size of the substudies. The CAN-BIND program includes a multimodal imaging database supported by extensive clinical, demographic, neuropsychological and biological data from people with major depression. It is a resource for Canadian investigators who are interested in understanding whether aspects of neuroimaging — alone or in combination with other variables — can predict the outcomes of various treatment modalities.
Introduction
Treatment of major depressive disorder (MDD) is evidence-based, but treatment selection is not personalized to the features of an individual’s illness.1 The discovery of biomarkers — or predictors — of treatment response is a priority in MDD research.2 A major challenge for identifying patient characteristics that predict treatment response is that MDD is a complex, heterogeneous condition. Current diagnostic systems codify depressive symptoms as criteria for MDD,3 but these symptoms are not unique to depression and, even if clustered together, they may not represent a single underlying disease process or treatment substrate.
A growing number of clinical studies are using MRI in an attempt to identify biomarkers of disease (for example, Jack and colleagues4), including depression (see Fonseka and colleagues5 for a recent review of studies using MRI to define markers of outcome in MDD). One approach to the detection of imaging biomarkers is to integrate data from large numbers of patients collected in independent studies. Keshavan and colleagues6 examined the circumstances under which a study could forgo efforts at protocol harmonization and phantom-based correction, relying only on the power of the data. They performed a scan–rescan study on 20 scanners with similar but nonidentical imaging parameters and determined that, in the absence of protocol harmonization, the sample size required could be in the thousands. The Enhancing NeuroImaging Genetics through Meta-Analysis (ENIGMA) consortium is a collaborative network of researchers who have integrated primarily structural data from more than 12 000 participants and 70 institutions around the world.7 The ENIGMA consortium has a working group focused on MDD that has reported on both subcortical8 and cortical brain structures.9 However, despite the power of this approach to examine factors such as age of onset and recurrence, ENIGMA’s psychiatric cohorts vary in terms of inclusion and exclusion criteria, duration of illness, the absence or presence of comorbid conditions, treatment history, ethnicity and other factors, limiting investigators’ ability to examine imaging data in the context of relevant clinical variables.9
An alternative approach to combining data from multiple independent studies is to conduct coordinated, multisite imaging studies. Several consortia have established guidelines and protocols for such studies, including the Function Biomedical Informatics Research Network (fBIRN),10 the Alzheimer’s Disease Neuroimaging Initiative (ADNI),4,11 the Mind Clinical Imaging Consortium (MCIC),12 the North American Imaging in Multiple Sclerosis (NAIMS) Cooperative13 and the Ontario Neurodegenerative Disease Research Initiative (ONDRI).14 However, only a few studies to date have employed multimodal, multisite imaging analyses to predict treatment outcomes in MDD.
The international Study to Predict Optimized Treatment in Depression (iSPOT)15 enrolled more than 2000 patients with MDD across 20 sites, but they recruited only 10% of the participants into the neuroimaging substudy, which was conducted at 2 sites.15,16 The iSPOT neuroimaging protocol included high-resolution 3-dimensional T1-weighted scans; diffusion tensor imaging (DTI); and T2-weighted proton density scans, as well as task-based functional MRI (fMRI) sequences to assess cognitive and emotional processing.16 The Establishing Moderators and Biosignatures of Antidepressant Response in Clinical Care (EMBARC) study17 enrolled 309 patients with early-onset MDD across 6 sites. The EMBARC neuroimaging protocol included 3-dimensional T1-weighted scans, DTI, arterial spin labelling and task-based fMRI sequences to assess the processing of reward and emotional conflict.
The Canadian Biomarker Integration Network in Depression (CAN-BIND; www.canbind.ca; see Kennedy and colleages18 and Lam and colleagues19) is a national program in depression research, funded by the Ontario Brain Institute, that seeks to address remaining gaps in the literature on response prediction by scanning approximately 1000 patients with depression or risk for depression. The CAN-BIND program includes multiple projects and has recruited approximately 1300 participants to date, including about 1000 with depression and 300 healthy participants for comparison. The CAN-BIND neuroimaging platform relies on evidence that data from different scanners are sufficiently robust to provide comparable results across multiple sites.20–23 Below, we briefly outline the substudies that use CAN-BIND imaging protocols.
The CAN-BIND-1 study includes 211 patients with MDD and 112 healthy controls. Medication-free patients were treated in an open trial protocol for 8 weeks with escitalopram, a selective serotonin reuptake inhibitor (SSRI). Nonresponders then had aripiprazole (an atypical antipsychotic) added to their regimen, and responders continued with escitalopram monotherapy for an additional 8 weeks (see Lam and colleagues19 for a detailed description). The study included MRI at baseline and after 2 and 8 weeks of treatment. It recruited participants from 6 sites in Canada (ClinicalTrials.gov identifier NCT01655706).
The CAN-BIND-2 study (Canadian rTMS Treatment and Biomarker Network in Depression; CARTBIND) explored the use of repetitive transcranial magnetic stimulation (rTMS), a noninvasive brain stimulation technique approved as a treatment for MDD. The CARTBIND trial is a 3-site study that uses 6 weeks of left dorsolateral prefrontal cortex intermittent theta-burst rTMS in patients with MDD, with the aim of identifying biomarkers of response to rTMS treatment. Scans have been obtained for 205 patients at baseline and within 1 week of completing rTMS therapy (ClinicalTrials.gov identifier NCT02729792).
The CAN-BIND-3 study (Canadian Psychiatric Risk and Outcome Study; PROCAN) is a 2-site study with the goal of improving the ability to identify youth at risk of serious mental illness, including MDD.24 In this study, 240 youth have been recruited, aged 12 to 25 years and at various levels of risk as defined in clinical staging models25 (e.g., genetic risk only, mild and/or attenuated symptoms, more pronounced but subthreshold symptoms). Participants are scanned at baseline and at 1- and 2-year follow-up, or when symptoms worsen.
The CAN-BIND-4 (Stress and Reward Anhedonia; SARA) single-site study aims to examine stress reactivity and reward responsivity as correlated domains of functioning in depression in 200 participants (100 patients with MDD, 100 healthy controls). Structural and functional brain imaging is being obtained at baseline and 6-month follow-up.
The CAN-BIND-5 (Biomarkers of Suicidality) single-site study has the goal of identifying an integrated biological marker model to predict risk of suicide attempt in MDD, and to test the stability of this model over time. Ninety patients with MDD with and without a history of suicide attempt, as well as 30 healthy controls, are being scanned at a baseline visit and at 1-year follow-up (ClinicalTrials.gov identifier NCT02811198).
The CAN-BIND-9 (Remote Cognitive Remediation for Depression; ReCoRD) single-site study aims to assess the effectiveness of cognitive remediation therapy in 75 participants with MDD who complete computer treatment modules from their homes. Participants are scanned at baseline and after online cognitive remediation, at 12- and 24-week follow-up.
The CAN-BIND-10 (Concussion and Depression Study) single-site study aims to characterize the biological profile of people with mild traumatic brain injury and depression, and to identify factors that may predict risk of depression after injury. Overall, 100 patients and 25 healthy controls are being scanned at entry into the study.
CAN-BIND participants
Participants are being recruited at 7 Canadian clinical centres: the University Health Network, the Centre for Addiction and Mental Health and Sunnybrook Health Sciences Centre in Toronto, Ontario; St. Joseph’s Healthcare in Hamilton, Ontario; Providence Care Hospital in Kingston, Ontario; Djavad Mowafaghian Centre for Brain Health in Vancouver, British Columbia; and the Mathison Centre for Mental Health Research and Education in the Hotchkiss Brain Institute, Calgary, Alberta. Each site has entered a standardized participation agreement with the Ontario Brain Institute to facilitate the transfer of both raw and processed/deidentified data, in accordance with the Ontario Brain Institute’s governance policy (www.braincode.ca/content/governance) and with any specific conditions required by each institution’s local legislative and ethical policies.
For all studies except CAN-BIND-3 and CAN-BIND-10, patients have a primary diagnosis of MDD, based on structured clinical interview. The CAN-BIND-324 study includes youth aged 12 to 15 years at varying degrees of risk for serious mental illness as defined by a clinical staging model.25 The CAN-BIND-10 study is recruiting patients with traumatic brain injury only and patients with both traumatic brain injury and MDD.
Across all studies, healthy participants for comparison have no history of psychiatric illness or current psychiatric illness as assessed by structured interview. Both patients and healthy participants are excluded if they have an estimated IQ of less than 70 based on the North American Adult Reading Test26; neurologic disease; a history of skull fracture or a severe or disabling medical condition; or a contraindication for MRI. Complete inclusion and exclusion criteria are specific to the various substudies.
CAN-BIND imaging protocols
The CAN-BIND program includes multiple longitudinal studies that employ common neuroimaging elements. Some use additional tasks and modalities, as indicated by the nature of the study. For the main characteristics and protocols for each CAN-BIND study, see Table 1, Table 2 and Appendix 1, Table S1 and Table S2. available at jpn.ca/180036.
Overview of CAN-BIND studies highlighting common, standardized data elements
Detailed scan acquisition parameters for structural MRI sequences
The CAN-BIND protocols include the following imaging sequences: a high-resolution 3-dimensional isotropic T1-weighted scan to assess fine anatomical detail and map cortical thickness; DTI to assess microstructural and white-matter integrity; and resting-state and task-based blood-oxygenation-level-dependent fMRI sequences to assess functional networks and pathways. The CAN-BIND-3 study also uses arterial spin labelling to measure cerebral blood flow. Protocols have been informed by a review of the relevant literature, consultation with other experts in the field and group consensus, taking into account each scanner’s capabilities.
Six scanner models are used across the clinical sites, mandating extensive and ongoing quality-control processes:21 a Discovery MR750 3.0 T (GE Healthcare), a Signa HDxt 3.0 T (GE Healthcare), a MAGNETOM Trio (Siemens Healthcare), a MAGNETOM Skyra (Siemens Healthcare), an Achieva 3.0 T (Philips Healthcare) and an Intera 3.0 T (Philips Healthcare).
Stimulus sizes, instructions to participants and support materials are standardized across sites. All behavioural data are captured using E-Prime version 2.0 Professional (Psychology Software Tools). For CAN-BIND-5 and CAN-BIND-10, PsychoPy,27 Inquisit (Millisecond) and Presentation (www.neurobs.com/) are also used. Guidelines and practices have been established for instructing participants to remain still throughout the scan, for applying a fiducial marker on the right temple, and for collecting respiratory bellows and peripheral gating (pulse oximetry) data using standard instruments provided by each manufacturer.
Whole-brain T1-weighted structural scan
Whole-brain T1-weighted structural scans are noninvasive, readily acquired and, because they are relatively short, generally well tolerated; these are features that may be important for identifying a potential biomarker.28 Structural MRI studies in patients with MDD have revealed widespread corticolimbic differences in grey matter29,30 and white matter,31 suggesting that there are detectable alterations in the structure of key brain regions that could inform clinically relevant outcomes. Studies examining how well structural MRI data may be able to diagnose depression report accuracy rates of 48% to 91%.32–37 Some studies have reported that structural alterations predict outcomes of treatment at the group level.38–46
The T1-weighted scans are acquired with a 3D isotropic resolution of 1 mm. For further detail on whole-brain T1-weighted imaging parameters, see Table 2. Information to confirm participant orientation is collected by placing a small vitamin E capsule on the right temple as a stereotactic marker (https://adni.loni.usc.edu/wp-content/uploads/2010/09/ADNI_MRI_Tech_Proc_Manual.pdf). Further information is included in Setup and Quality Assurance of MRI Protocols.
Whole-brain DTI
Diffusion tensor imaging studies have demonstrated altered white-matter microstructural abnormalities in patients with MDD. Decreased fractional anisotropy, a proxy measure of the directionality of diffusion, has been reported in patients with MDD in the frontal and occipital (fusiform) regions.47–50 Fibre tracking has revealed the involvement of similar structures in MDD.47 White-matter alterations have predicted treatment outcomes with up to 65% accuracy.33,37 In another study, elevated baseline fractional anisotropy in tracts connecting to the right amygdala has been associated with remission following SSRI treatment.51
The CAN-BIND DTI acquisition protocol employs a single-shot, spin-echo, echo planar imaging sequence with diffusion sensitizing gradients applied in 31 noncollinear directions (b = 1000 s/mm2) and 6 volumes with b = 0 s/mm2. For CAN-BIND-3, diffusion sensitizing gradients were applied in 45 noncollinear directions, with 8 images collected at b = 1000 s/mm2 and 8 images collected at b = 2500 s/mm2. Increasing the number of diffusion-encoded directions improves the accuracy and/or robustness of diffusion tensor estimation,52 and having more directions allows for the removal of any corrupted directions (e.g., due to motion/movement).53 See Table 2 for further details on the parameters for whole-brain DTI.
Resting-state fMRI
Resting-state fMRI allows for the identification of task-independent and spontaneous neural activation that coincides temporally to form neural networks54 such as the default mode network (e.g., see Greicius and colleagues55), the salience network or cognitive control network (e.g., Menon, 56 Menon and Uddin,57 or Seeley and colleagues58), and the affective network.59–63 The default mode network shows abnormal patterns of functional connectivity in MDD55,64–66 that may normalize following treatment67,68 or may be associated with treatment resistance.69
Resting-state data are collected over a 10-minute scan during which participants are instructed to lie still, keep their eyes open and focus on a fixation cross.70 Standardized instructions are used across sites. Images are obtained using a whole-brain T2*-sensitive blood-oxygen-level-dependent echo planar imaging series, with a repetition time of 2000 ms, an echo time of 30 ms and voxel dimensions of 4 mm × 4 mm × 4 mm, kept constant across sites and scanners. See Table 3 for further details on the parameters for resting-state fMRI.
Detailed scan acquisition parameters for resting-state functional MRI sequences
Task-based fMRI
Task-based fMRI studies suggest that there may be different patterns of change associated with specific treatments or classes of treatment.68,71–76 The CAN-BIND substudies test treatment- and population-specific questions, using cognitive-functional tasks that are described in detail in Appendix 1.
Task-relevant instructions are standardized and given before the scan sessions. Each site uses a comparable, custom-manufactured, magnet-compatible input device (www.mrn.org/collaborate/imaging-equipment) to record participants’ responses. Acquisition parameters are similar to those for resting-state fMRI, and are listed in detail in Appendix 1, Table S1 and Table S2.
Arterial spin labelling
Arterial spin labelling perfusion MRI measures regional cerebral blood flow and may be used to study subtle brain perfusion changes in psychiatric illnesses. Perfusion patterns may hold promise as objective biomarkers for tracking illness progression, as well as pharmacological/treatment effects in various neuropsychiatric disorders.77
Data storage
Clinical data are collected and stored in the Ontario Brain Institute’s Centre for Ontario Data Exploration (Brain-CODE; www.braincode.ca/; Vaccarino and colleagues78). This online neuroinformatics platform allows researchers to collaborate across distances and work efficiently at multiple sites. Brain-CODE is deployed at the Centre for Advanced Computing at Queen’s University in Kingston, Ontario. The Centre for Advanced Computing is a member of the Compute Canada high-performance computing consortium, which supports regulatory-compliant processes for securing the privacy of health care data (https://cac.queensu.ca/overview). Online clinical and neuroimaging data are accessed on secure websites via restricted portals that require unique usernames and passwords for each member of the study team. User profiles are assigned only to study personnel who require access to enter and verify data, and credentials for each user are vetted by the program manager.
The SPReD database (originally the Stroke Patient Recovery Research Database) is a comprehensive online repository powered by the open-source Extensible Neuroimaging Archiving Toolkit (XNAT) imaging informatics platform, 79,80 where neuroimaging data are uploaded and stored. Structural and functional MRI data are uploaded from each site as Digital Imaging and Communications in Medicine (DICOM) images. Supplementary records, such as behavioural and physiological data, and session notes associated with an imaging session, are uploaded through a special subprocess.
Neuroinformatics framework
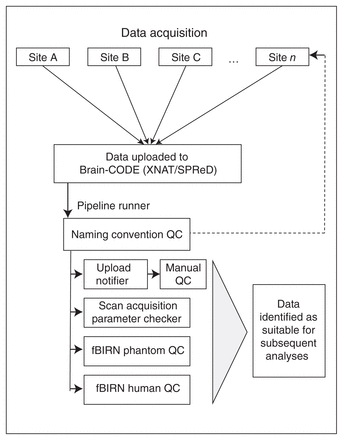
The CAN-BIND neuroinformatics framework consists of software, tools, pipelines and procedures designed to ensure high-quality data acquisition, databasing, archiving, assessment, analysis and tracking, an overview of which is shown in Figure 1. The primary platform for this set of tools is XNAT/SPReD, provided through Brain-CODE. In addition to the MRI data being captured and managed through XNAT/SPReD, other study-related data are captured using OpenClinica and RedCap. A visualization “dashboard” built using SpotFire (http://spotfire.tibco.com/) is used to upload aggregated data tracking and analytics results from phantom data (see Fig. 2 and Fig. 3).
Overview of the CAN-BIND neuroinformatics framework. Data from each site is uploaded to Brain-CODE, where specifically designed pipelines check the data for compliance with scan acquisition parameters, naming convention and completeness. Automatic messages are sent to initiate manual QC. The CAN-BIND neuroinformatics framework also includes pipelines for the analysis of phantom data. CAN-BIND = Canadian Biomarker Integration Network in Depression; fBIRN = Functional Biomedical Informatics Research Network; QC = quality control; SPReD = originally named the Stroke Patient Recovery Research Database; XNAT = Extensible Neuroimaging Archiving Toolkit.
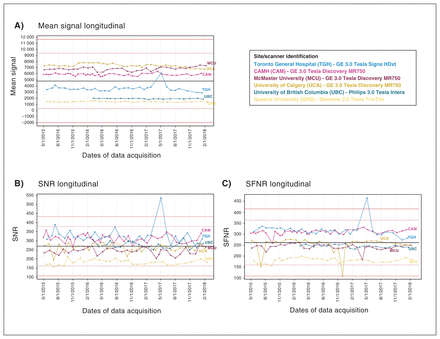
Examples of data quality tracking and assessment pipelines. Phantom data are tracked longitudinally to monitor adherence and data quality of imaging protocols. Illustrated here is an example where spiking in the overall mean signal intensity across acquired images at one data acquisition site (light blue) was tracked to be related to its SNR and its SNFR. (A) Mean signal longitudinal: this metric tracks the average overall signal intensity across all voxels and images, per scanning session. (B) SNR longitudinal: this metric tracks the average overall SNR. The mean SNR is the static spatial noise × image across a 21 × 21 voxel region of interest centred on the image. The signal summary value is the average of the signal image across this same region of interest. Then, SNR = (signal summary value)/√(variance summary value/number of time points). (C) SFNR longitudinal: the SFNR is the voxel-wise ratio of the temporal variance standard deviation and temporal mean intensity of the 4-dimensional phantom image after quadratic detrending. The SFNR summary value is the mean SFNR value within the evaluation region of interest (a 21 × 21 voxel region in the centre of the image). SFNR = signal-to-fluctuation-noise ratio; SNR = signal-to-noise ratio.
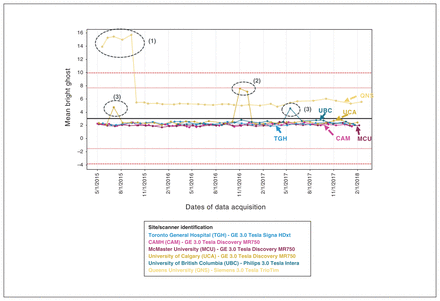
Examples of data quality tracking and assessment pipelines. Phantom data are tracked longitudinally to monitor adherence and data quality of imaging protocols. Illustrated here are examples where the mean intensity of ghost-only voxels showed deviations; investigation and explanation of these anomalies are listed in 1, 2 and 3, below. Mean bright ghost longitudinal: ghost metrics are calculated for each volume by taking a dilated mask (“original mask”) of the data, and shifting it by N/2 voxels in the appropriate axis to create a “ghost mask.” Whereas the mean intensities of those voxels in the ghost mask and not in the original mask is the “mean ghost” value, the “mean bright ghost” is the mean intensity of the top 10% of ghost-only voxels. (1) Anomaly: investigation led to protocol adjustments. (2) Receiver coil failure: addressing failure resulted in data returning to the level seen previously. (3) Anomalies, investigation: corresponding human functional MRI scans acquired around this date appeared fine; subsequent phantom scans were fine.
CAN-BIND quality control and quality assurance procedures
The importance of quality assurance and control in multisite studies is recognized.81 The full spectrum of data quality control and data quality assurance methods was implemented early in CAN-BIND-1. These methods are described in the sections that follow and have been applied to most of the CAN-BIND substudies. The CAN-BIND-2 and CAN-BIND-3 studies have not been uploading their data to SPReD, so the automated adherence checks described here do not apply to them.
Quality control
Data file-naming convention and adherence checks
Participants are assigned unique identification codes, which are standardized to contain a program code (3 letters), a study number (2 digits), a site identification code (3 letters) and a participant number (4 digits; e.g., CBN01_UCA_0001). These file-naming conventions are applied to MRI and behavioural data files. A pipeline assessing the consistency of naming conventions is implemented in XNAT/SPReD; if noncompliance is detected, notification is sent to relevant study personnel asking them to implement corrections, with follow-up until corrections are performed. The data will not undergo subsequent quality-control checks until file-naming conventions have been adhered to.
Parameter adherence checks of MRI protocols
Also implemented in SPReD is a quality-control pipeline for MRI protocols, which compares the acquisition parameters of newly uploaded scans against a reference protocol. Reference protocols have been established for each site and scanner type, taking into account the fact that scan parameters are necessarily different among scanners and manufacturers. The reference protocol defines the sequences and appropriate acquisition parameters (values) for each sequence. If discrepancies are identified between the data uploaded and the reference protocol, e-mail notifications are sent to study personnel, asking them to identify causes for adherence check failures and pointing to the need for possible rescanning.
Image quality
It is necessary to obtain images of sufficient subjective quality, free of motion artifacts, covering a full field of view and free of other scanner-related artifacts in order to process the data through various pipelines. Certain sequences, such as resting-state fMRI, are more susceptible to motion and other artifacts. Others, such as T1-weighted images, are of such paramount importance that tolerance for motion or other artifacts is low because they influence the quality of the data and the usability of other sequences, which are typically coregistered to T1-weighted scans. Trained expert quality-control raters are automatically notified when new data are uploaded to SPReD. They perform visual assessment of the MRI data image quality using the SPReD interface. The quality-control raters have received training via ONDRI, based on the data quality control protocol from the Centre for Brain Science at Harvard University.82 Raters compare their assessments and comments on scan quality for subsets of data collected at participating CAN-BIND sites. Each imaging sequence is reviewed independently for quality, including full-brain coverage (on a 2-point scale: complete or incomplete), motion and other image artifacts (on a 3-point scale: none, mild or severe), based on the data quality-control protocol. 82 Imaging that has insufficient coverage, excessive motion as identified by visual inspection of rigid uniform stripes running horizontally across the brain82 or other imaging artifacts that may interfere with future processing and usability are marked as questionable or unusable, depending on severity. If images are flagged as unusable, they are unavailable for subsequent analysis, and a request is made to the study site to rescan the participant whenever feasible. An upload delay dashboard also serves to inform program managers of the delay time in uploading data once it has been acquired.
Assessment of site differences
Cross-site T1 piloting included a travelling participant or “ human phantom,” who travelled to each CAN-BIND-1 site for anatomic scans to document within and between-site variance.
Setup and quality assurance of MRI protocols
Setup of scan parameters
Prior to study launch, scan parameters from DICOM header files were examined to match scan parameters across CAN-BIND-1 sites to a majority consensus where possible (exceptions included receiver bandwidth of multiple scans and acceleration type). Quality assurance test sample data were acquired from CAN-BIND-1 sites and examined; then, recommendations were made to each site if adjustments or revisions were required. Acquisition parameters were transferred digitally from sites with a GE Discovery 750, and by PDF from sites with other scanners. Technologists entered the values and subsequently checked the DICOM headers. Following revisions, this process was repeated. Next, a human phantom/expert visited the CAN-BIND-1 sites to identify scanning issues and acquire data. Subsequently, T1 acquisitions at sites without GE Discovery 750 scanners were matched by acquiring multiple flip-angle/inversion time values and identifying those that gave a contrast-to-noise measure that was most similar to the GE Discovery 750 scanners. The T1 scans were prescribed to a sagittal acquisition using a nonselective radiofrequency pulse; fMRI and DTI scans were acquired as true axial scans to reduce cross-site variation. In addition, fMRI and DTI scans were acquired using fat saturation at all sites, rather than having some sites use spectral–spatial pulses. Protocol corrections were made to ensure that image resampling after acquisition was not performed (rhimsize was set on GE scanners to prevent interpolation of images in DTI). Acquisition in DTI was shortened from a repetition time of 14 000 ms to one of 9000 ms. This reduced the scan time, while leaving contrast unaffected, because it maintained at least 6 T1 time constants for both grey and white matter. Scan acquisition parameters according to site for structural MRI data — including 3-dimensional T1-weighted scans, DTI scans and T2-weighted proton density scans — are listed in Table 2, and parameters for functional MRI data are listed in Table 3 and Appendix 1, Table S1 and Table S2.
Monitoring and quality assurance of imaging parameters
Since the CAN-BIND launch, all CAN-BIND-1 sites have obtained monthly scans of 2 geometric phantoms (a spherical agar phantom developed by the fBIRN consortium, and a custom-built cylindrical model using plastic LEGO blocks)20,83 to facilitate scanner calibration and troubleshooting over the long term. Examples of the longitudinal tracking of phantom data quality are illustrated in Figure 2 and Figure 3. Phantom scans are also acquired at St. Michael’s Hospital for CAN-BIND-5 and CAN-BIND-10. Phantom scans are not collected at Sunnybrook Health Sciences Centre, the second data acquisition site for CAN-BIND-3.
Setup of fMRI paradigms
To standardize the viewing angle for fMRI task stimuli, a standard grid was displayed at each site, viewing distance was measured, and the visual angle of the projected image was calculated. Consistent cross-site viewing angle was established using specific display parameters in the E-Prime software for each site. Across sites, the version of the E-Prime stimulus display software was matched. Button responses and ASCII key codes were confirmed and used in site-specific E-Prime task versions. Data files produced by each paradigm were examined to confirm that the proper response information was being acquired and logged.
Sites were also provided with a scripted set of instructions to be issued before resting-state scans, as well as a standardized fixation cross for participants to focus on during the resting state scan. A set of participant orientation/training slides were instituted for functional tasks. Randomization schedules were provided for the functional task version administered (e.g., A/B/C for the go/no-go task) and task order between, for example, go/no-go and reward tasks. For detail on fMRI tasks, see Appendix 1. Study coordinators were provided with a guide to follow when checking the fidelity of the acquired behavioural data. Finally, conference calls were held with the research coordinators at each site to ensure that standard operating procedures were communicated and instituted.
Discussion
The neuroinformatics procedures and pipelines employed in CAN-BIND address many challenges associated with combining MRI data from multisite studies. Considerable effort has been focused on the image acquisition protocols, and procedures have been implemented — automated, where possible — to ensure the ongoing quality of the images. We recognize, however, that residual differences in neuroimaging data collected across different sites and MRI vendors will likely still exist.
The “reproducibility in science crisis”84 has required that imaging studies examine common approaches to study design, monitoring and interpretation. Issues underlying the difficulty with replication are multifaceted, and protocols are emerging to ensure that imaging studies are well planned, well executed and well reported. This includes making the details of how studies are designed, executed and analyzed more apparent and transparent.85,86 This paper aims to provide methodological detail for the CAN-BIND studies in a transparent and comprehensive manner. As evident from Figure 1, there are common data elements across the CAN-BIND program substudies, specifically for 3D anatomic scans, resting-state fMRI and DTI. Scan parameters (as detailed in Table 2 and Table 3) are as comparable and compatible as scanner manufacturer and type allow. Quality-control procedures, such as checking protocol adherence for participant scans and manual quality control of acquired data, are performed for most substudies, based on an agreed-upon protocol. For example, although CAN-BIND-2 and CAN-BIND-3 are not currently uploading data to SPReD for automatic protocol adherence checks, data are being manually inspected for data quality.
Limitations
Although we consider it a strength that the CAN-BIND protocol is applied across participants with a wide age range (12 to 70 years), age-related differences will need to be assessed with caution, as will differences in sex and other demographic factors. We did not assess for the presence of cerebrovascular disease in our sample, although there is an association between cardiovascular disease and MDD,87 but also with MDD and other medical conditions.88 Given the relatively young age of our samples (e.g., Lam and colleagues,19 Addington and colleagues,24 Santesteban-Echarri and colleages89 and Kennedy and colleagues90), this is unlikely to be a driving factor in neuroimaging results, but medical comorbidity is an important consideration in studies of psychiatric disease. No routine screening for substance use was performed, potentially affecting our findings. As noted above, CAN-BIND-2 and CAN-BIND-3 are not subject to the automatic adherence checks that would result from uploading to SPReD.
Conclusion
The CAN-BIND program is unusual in that it uses a suite of common imaging protocols across a variety of studies that examine predictive markers of response to various treatment modalities in MDD. Although each CAN-BIND substudy is expected to yield valuable information, the consistent protocols, centralized data collection and quality control that will eventually allow for cross-study investigations is likely to be the greatest strength of CAN-BIND.
Deidentified CAN-BIND data eventually will be shared by the Ontario Brain Institute with other collaborators and third parties for research purposes.91 These data sets may inform clinical research teams with similar data sets comparing MDD with other psychiatric conditions, or comparing different treatment modalities. Thus, rigorous, recorded quality control of CAN-BIND neuroimaging and related data are crucial for ensuring the value of this data set to the greatest number of investigators. When fully realized, the CAN-BIND data set will provide a comprehensive resource for researchers interested in predictors, moderators and mediators of response to treatment in MDD.
Acknowledgements
CAN-BIND is an Integrated Discovery Program carried out in partnership with, and with financial support from, the Ontario Brain Institute, an independent nonprofit corporation funded partially by the Ontario government. The opinions, results and conclusions are those of the authors, and no endorsement by the Ontario Brain Institute is intended or should be inferred. Additional funding is provided by the Canadian Institutes of Health Research, Lundbeck, Bristol-Myers Squibb and Servier. Funding and/or in-kind support is also provided by the investigators’ universities and academic institutions. All study medications are independently purchased at wholesale market values.
Footnotes
↵* These authors share first authorship.
Competing interests: G. MacQueen reports consultancy/speaker fees from Lundbeck, Pfizer, Johnson & Johnson and Janssen, outside the submitted work. B. Frey reports grants and personal fees from Pfizer and personal fees from Sunovion, outside the submitted work. R. Milev reports grants, nonfinancial support and honoraria from Lundbeck, Janssen and Pfizer; personal fees and honoraria from Sunovion, Shire, Allergan and Otsuka; grants from Boehringer Ingelheim; and grants from the Ontario Brain Institute, the Canadian Institutes for Health Research and CAN-BIND, outside the submitted work. F. Vila-Rodriguez reports nonfinancial support from Magventure during the conduct of the study; grants from the Canadian Institutes for Health Research, Brain Canada, the Michael Smith Foundation for Health Research, and the Vancouver Coastal Health Research Institute; and personal fees from Janssen, outside the submitted work. S. Rizvi reports grants from Pfizer Canada, outside the submitted work. S. Strother reports grants from Canadian Biomarker Integration Network in Depression during the conduct of the study and grants from Ontario Brain Institute, outside the submitted work. He is also the chief scientific officer of the neuroimaging data analysis company ADMdx, Inc (www.admdx.com), which specializes in brain image analysis to enable diagnosis, prognosis and drug effect detection for Alzheimer disease and various other forms of dementia. R. Lam reports grants from Canadian Institutes of Health Research during the conduct of the study; grants from Asia-Pacific Economic Cooperation, VGH-UBCH Foundation, BC LEading Edge Endowment Fund, Janssen, Lundbeck, Pfizer and St. Jude Medical, outside the submitted work; personal fees from Allergan, Akili, CME Institute, Canadian Network for Mood and Anxiety Treatments, Janssen, Lundbeck, Lundbeck Institute, Pfizer, Otsuka, Medscape and Hansoh, outside the submitted work; travel expenses from Asia-Pacific Economic Cooperation outside the submitted work; and stock options from Mind Mental Health Technologies.
Contributors: G. MacQueen, S. Hassel, J. Addington, C. Bowie, S. Bray, J. Downar, J. Foster, B. Frey, B. Goldstein, K. Harkness, C. Lebel, R. Milev, D. Müller, S. Parikh, S. Rizvi, S. Rotzinger, C. Soares, S. Strother and S. Kennedy conceived and designed the article. R. Lam acquired the data. S. Hassel, S. Arnott, A. Davis, J. Harris, G. Sharma, J. Yu, M. Zamyadi and S. Strother analyzed and interpreted the data. G. MacQueen and S. Hassel wrote the article, which all authors reviewed. All authors approved the final version to be published and can certify that no other individuals not listed as authors have made substantial contributions to the paper.
Members of the CAN-BIND Investigator Team: www.canbind.ca/our-team
- Received March 11, 2018.
- Revision received July 23, 2018.
- Revision received October 15, 2018.
- Accepted October 18, 2018.